publications
publications by categories in reversed chronological order, generated by jekyll-scholar.
| Google Scholar profile ORCiD record |    |
2026
-
 Performance Evaluation of Large Language Models in Multilingual Medical Multiple-Choice Questions: Mixed Methods StudyLivia Maria Strasser, Wilma Anschuetz, Fabio Dennstädt, and Janna HastingsJMIR Medical Education, Mar 2026
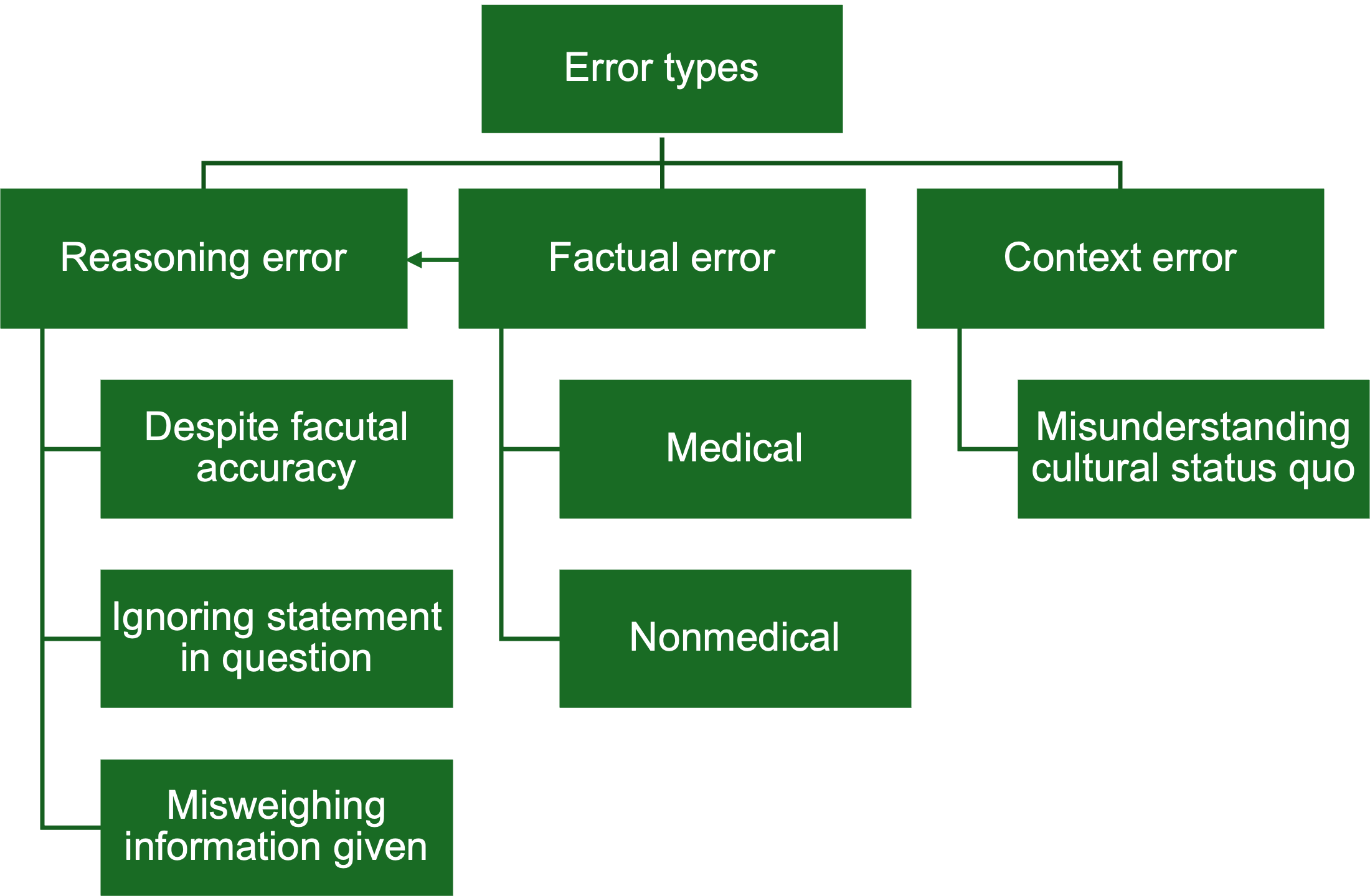
Performance Evaluation of Large Language Models in Multilingual Medical Multiple-Choice Questions: Mixed Methods StudyLivia Maria Strasser, Wilma Anschuetz, Fabio Dennstädt, and Janna HastingsJMIR Medical Education, Mar 2026Background: Artificial intelligence continues to transform health care, offering promising applications in clinical practice and medical education. While large language models (LLMs), as a form of generative artificial intelligence, have shown potential to match or surpass medical students in licensing examinations, their performance varies across languages. Recent studies highlight the complex influence and interdependency of factors such as language and model type on LLMs’ accuracy; yet, cross-language comparisons remain underexplored. Objective: This study evaluates the performance of LLMs in answering medical multiple-choice questions quantitatively and qualitatively across 3 languages (German, French, and Italian), aiming to uncover model capabilities in a multilingual medical education context. Methods: For this mixed methods study, 114 publicly accessible multiple-choice questions in German, French, and Italian from an online self-assessment tool were analyzed. A quantitative performance analysis of several LLMs developed by OpenAI, Meta AI, Anthropic, and DeepSeek was conducted to evaluate their performance on answering the questions in text-only format. For the comparative analysis, a variation of input question language (German, French, and Italian) and prompt language (English vs language-matched) was used. The 2 best-performing LLMs were then prompted to provide answer explanations for incorrectly answered questions. A subsequent qualitative analysis was conducted on these explanations to identify the reasons leading to the incorrect answers. Results: The performance of LLMs in answering medical multiple-choice questions varied by model and language, showing substantial differences in accuracy (between 64% and 87%). The effect of input question language was significant (\textless.01) with models performing best on German questions. Across the analyzed LLMs, prompting in English generally led to better performance in comparison to language-matched prompts, but the top-performing models exceptionally showed comparable results for language-matched prompts. Qualitative analysis revealed that answer explanations of the analyzed models (GPT4o and Claude-Sonnet-3.7) showed different reasoning errors. In several explanations, this occurred despite factual accuracy on the represented topic. Furthermore, this analysis revealed 3 questions to be insufficiently precise. Conclusions: Our results underline the potential of LLMs in answering medical examination questions and highlight the importance of careful consideration of model choice, prompt, and input languages, because of relevant performance variability across these factors. Analysis of answer explanations demonstrates a valuable use case of LLMs for improving examination question quality in medical education, if data security regulations permit their use. Human oversight of language-sensitive or clinically nuanced content remains essential to determine whether incorrect output stems from flaws in the questions themselves or from errors generated by the LLMs. There is a need for ongoing evaluation as well as transparent reporting to ensure reliable integration of LLMs into medical education contexts.
2025
-
 Automated gene identification in oncology literature: A comparative evaluation of Natural Language Processing approachesMarie Wosny and Janna HastingsStudies in Health Technology and Informatics, Jul 2025
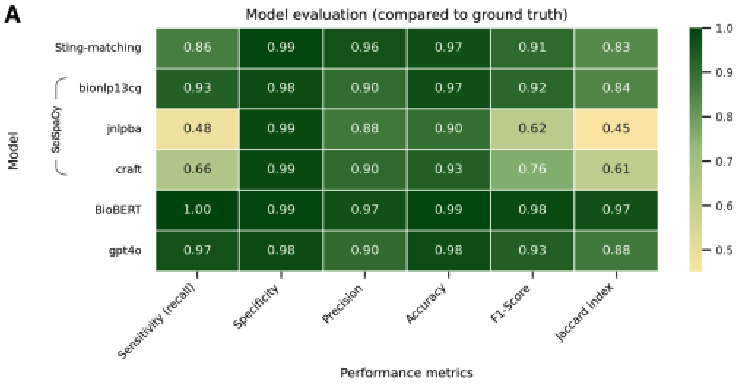
Automated gene identification in oncology literature: A comparative evaluation of Natural Language Processing approachesMarie Wosny and Janna HastingsStudies in Health Technology and Informatics, Jul 2025The exponential growth of biomedical literature necessitates automated approaches for extracting biological entities, such as genes, to support research. This study systematically compares rule-based, Named Entity Recognition (NER)-based, and transformer-based models for extracting 161 Oncomine™ genes from 100 cancer-related abstracts. The transformer-based BioBERT model achieved the highest recall (1.00) and F1-score (0.98), followed by GPT-4o, which, despite its effectiveness, required substantial computational resources. NER-based scispaCy models exhibited varying performance, while rule-based string-matching demonstrated high precision but lower recall. The finding highlights the trade-offs between accuracy and computational efficiency, emphasizing the potential for hybrid approaches in large-scale text mining applications.
-
 Large Language Models for Detection of Genetic Variants in Biomedical LiteratureMarie Wosny and Janna HastingsStudies in Health Technology and Informatics, Aug 2025
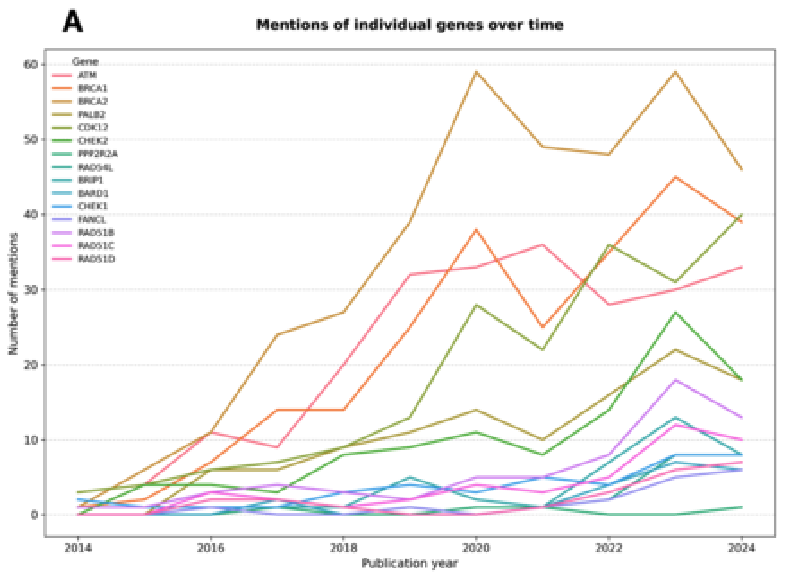
Large Language Models for Detection of Genetic Variants in Biomedical LiteratureMarie Wosny and Janna HastingsStudies in Health Technology and Informatics, Aug 2025Personalized medicine relies on knowledge about genetic variants, yet clinicians face challenges in keeping track of these throughout the large volume of literature. We applied a Large Language Model (LLM) to analyze prostate cancerrelated literature and detect variant mentions. Among 18,936 publications analyzed, 797 referenced relevant genes, of which 23 contained genetic variant mentions. The LLM successfully identified all 23 relevant publications, with only six false positives, resulting in a precision of 79.31%, a perfect recall of 100%, and an overall accuracy of 99.25%. Moreover, the LLM inferred additional details not explicitly stated in the articles, enriching the knowledge base. These findings highlight the potential of LLMs to augment human expertise and improve efficiency in literature review. Nevertheless, rigorous validation and oversight remain critical to address limitations and ensure careful integration into clinical workflows.
-
 A CDE-based data structure for radiotherapeutic decision-making in breast cancerFabio Dennstädt, Maximilian Schmalfuss, Johannes Zink, Janna Hastings, Roberto Gaio, Max Schmerder, Paul Martin Putora, and Nikola CihoricBMC Medical Informatics and Decision Making, Feb 2025
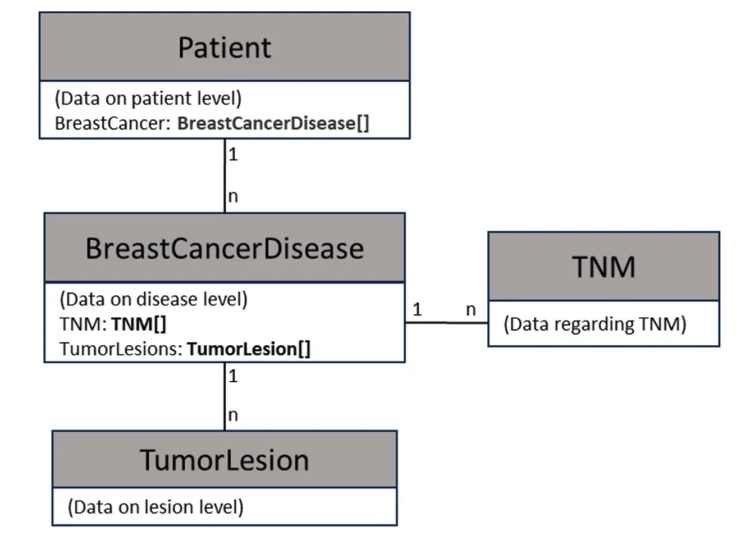
A CDE-based data structure for radiotherapeutic decision-making in breast cancerFabio Dennstädt, Maximilian Schmalfuss, Johannes Zink, Janna Hastings, Roberto Gaio, Max Schmerder, Paul Martin Putora, and Nikola CihoricBMC Medical Informatics and Decision Making, Feb 2025Background The increasing complexity and data-driven nature of oncology and radiation therapy necessitates structured and precise data management strategies. The National Institutes of Health (NIH) has introduced Common Data Elements (CDEs) as a uniform approach to facilitate consistent data collection. However, there is currently a lack of a comprehensive set of CDEs for describing situations for and within radiation oncology. Aim of this study was to create a CDE-based data structure for radiotherapeutic decision-making in breast cancer to promote structured data collection on the level of a local hospital. Methods Local Standard Operating Procedures (SOPs) were analyzed to identify relevant decision-making criteria used in clinical practice. Corresponding CDEs were identified and a structured data framework based on these CDEs was created. The framework was translated into machine-readable JSON format. Six clinical practice guidelines of the American Society for Radiation Oncology (ASTRO) were analyzed as full-text to investigate how many guideline recommendations and corresponding decision-making criteria could be presented using the data structure. Results The study identified 31 decision-making criteria mentioned in the SOPs, leading to the establishment of 46 CDEs. A hierarchical structure within an object-oriented data framework was created and converted into JSON format. 94 recommendations with mentioning of decision-making criteria in 216 cases were identified across the six ASTRO guidelines. In 151 cases (70.0%) the mentioned criterion could be presented with the data framework. Conclusions The CDE-based data structure represents a clear framework for structuring medical data for radiotherapeutic decision-making in breast cancer patients. The approach facilitates detailed description of individual breast cancer cases and aids in the integration of information technology. Furthermore, it promotes sharing of standardized data among healthcare providers.
- Finding Consensus on Trust in AI in Health Care: Recommendations From a Panel of International ExpertsGeorg Starke, Felix Gille, Alberto Termine, Yves Saint James Aquino, Ricardo Chavarriaga, Andrea Ferrario, Janna Hastings, Karin Jongsma, Philipp Kellmeyer, Bogdan Kulynych, and 8 more authorsJournal of Medical Internet Research, Feb 2025
Background: The integration of artificial intelligence (AI) into health care has become a crucial element in the digital transformation of health systems worldwide. Despite the potential benefits across diverse medical domains, a significant barrier to the successful adoption of AI systems in health care applications remains the prevailing low user trust in these technologies. Crucially, this challenge is exacerbated by the lack of consensus among experts from different disciplines on the definition of trust in AI within the health care sector. Objective: We aimed to provide the first consensus-based analysis of trust in AI in health care based on an interdisciplinary panel of experts from different domains. Our findings can be used to address the problem of defining trust in AI in health care applications, fostering the discussion of concrete real-world health care scenarios in which humans interact with AI systems explicitly. Methods: We used a combination of framework analysis and a 3-step consensus process involving 18 international experts from the fields of computer science, medicine, philosophy of technology, ethics, and social sciences. Our process consisted of a synchronous phase during an expert workshop where we discussed the notion of trust in AI in health care applications, defined an initial framework of important elements of trust to guide our analysis, and agreed on 5 case studies. This was followed by a 2-step iterative, asynchronous process in which the authors further developed, discussed, and refined notions of trust with respect to these specific cases. Results: Our consensus process identified key contextual factors of trust, namely, an AI system’s environment, the actors involved, and framing factors, and analyzed causes and effects of trust in AI in health care. Our findings revealed that certain factors were applicable across all discussed cases yet also pointed to the need for a fine-grained, multidisciplinary analysis bridging human-centered and technology-centered approaches. While regulatory boundaries and technological design features are critical to successful AI implementation in health care, ultimately, communication and positive lived experiences with AI systems will be at the forefront of user trust. Our expert consensus allowed us to formulate concrete recommendations for future research on trust in AI in health care applications. Conclusions: This paper advocates for a more refined and nuanced conceptual understanding of trust in the context of AI in health care. By synthesizing insights into commonalities and differences among specific case studies, this paper establishes a foundational basis for future debates and discussions on trusting AI in health care.
-
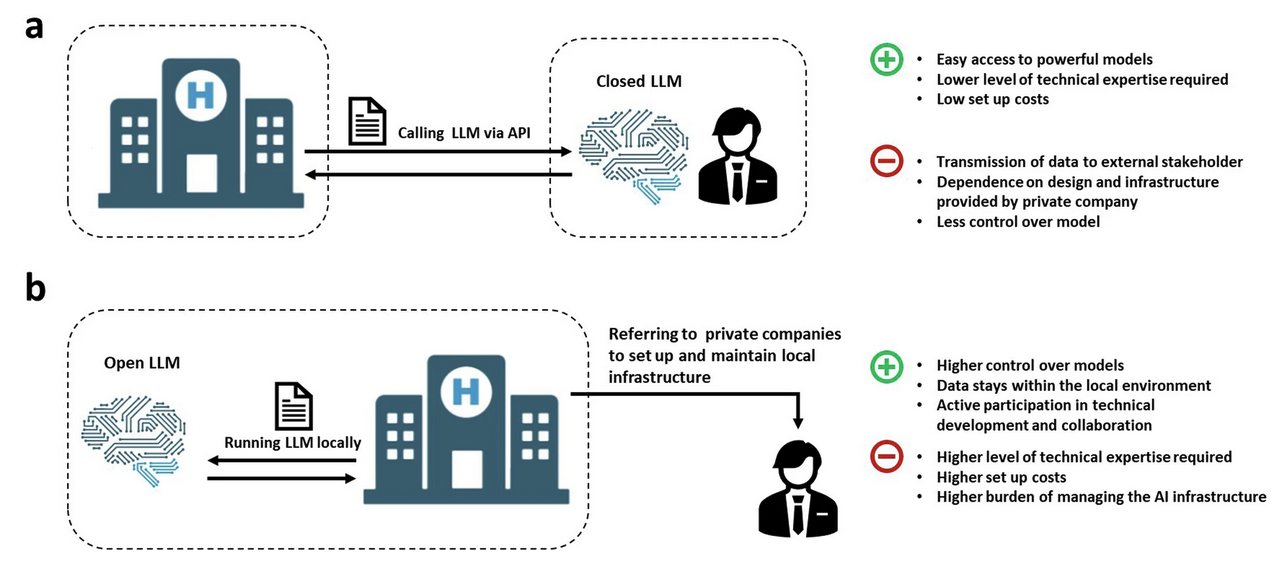 Implementing large language models in healthcare while balancing control, collaboration, costs and securityFabio Dennstädt, Janna Hastings, Paul Martin Putora, Max Schmerder, and Nikola Cihoricnpj Digital Medicine, Mar 2025
Implementing large language models in healthcare while balancing control, collaboration, costs and securityFabio Dennstädt, Janna Hastings, Paul Martin Putora, Max Schmerder, and Nikola Cihoricnpj Digital Medicine, Mar 2025Integrating Large Language Models (LLMs) into healthcare promises substantial advancements but requires careful consideration of technical, ethical, and regulatory challenges. Closed LLMs of private companies offer ease of deployment but pose risks related to data privacy and vendor dependence. Open LLMs deployed on local hardware enable greater model customization but demand resources and technical expertise. Balancing these approaches, with collaboration among clinicians, researchers, and companies is crucial to ensure effective, secure, and ethical implementation.
- Implementing a Resource-Light and Low-Code Large Language Model System for Information Extraction from Mammography Reports: A Case StudyFabio Dennstädt, Simon Fauser, Nikola Cihoric, Max Schmerder, Paolo Lombardo, Grazia Maria Cereghetti, Sandro von Däniken, Thomas Minder, Jaro Meyer, Lawrence Chiang, and 12 more authorsApr 2025
Background Large Language Models (LLMs) have been successfully used to extract structured data from free-text radiology reports. Most of current studies were conducted with private models accessed via Application Programming Interface (API). We aimed to evaluate the feasibility of using open-source LLMs, deployed on limited local hardware resources for extraction of structured information from free-text mammography reports, according to a Common Data Elements (CDE)-based framework. Methods Seventy-nine CDEs were defined by an interdisciplinary expert panel, reflecting real-world reporting practice. Sixty-one reports were classified by two independent researchers with 1533 classifications assigned to establish ground truth. Five different open-source LLMs deployable on a single GPU were used for data extraction using the general-classifier Python package. Extractions were performed for two different prompt approaches with classification metrics calculated overall and on subgroups. Additional analyses were conducted using thresholds for the relative probability of classifications. Results High inter-rater agreement was observed between manual classifiers (Cohen’s Kappa 0.83). Using default prompts, the LLMs achieved accuracies of 59.23–72.86%. Adapting prompts to better explain classification tasks improved performance for all models, with accuracies of 64.71–85.32%. Setting certainty thresholds further improved accuracies to \textgreater90% but reduced the coverage rate to \textless50%. Conclusion Locally deployed open-source LLMs can effectively extract information from mammography reports with good accuracy, addressing data privacy concerns while maintaining compatibility with limited computational resources. Prompt engineering substantially increases performance, highlighting the importance of optimization in clinical applications. Using a CDE-based framework provides clear semantics and structure, facilitating interoperability and consistent data extraction.
-
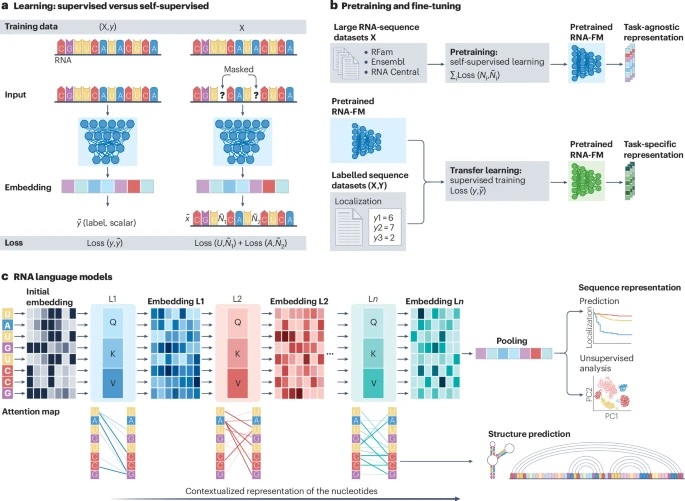 Decoding the interactions and functions of non-coding RNA with artificial intelligenceVincent Jung, Cédric Vincent-Cuaz, Charlotte Tumescheit, Lisa Fournier, Marousa Darsinou, Zhi Ming Xu, Ali Saadat, Yiran Wang, Petros Tsantoulis, Olivier Michielin, and 8 more authorsNature Reviews Molecular Cell Biology, Jun 2025
Decoding the interactions and functions of non-coding RNA with artificial intelligenceVincent Jung, Cédric Vincent-Cuaz, Charlotte Tumescheit, Lisa Fournier, Marousa Darsinou, Zhi Ming Xu, Ali Saadat, Yiran Wang, Petros Tsantoulis, Olivier Michielin, and 8 more authorsNature Reviews Molecular Cell Biology, Jun 2025In addition to encoding proteins, mRNAs have context-specific regulatory roles that contribute to many cellular processes. However, uncovering new mRNA functions is constrained by limitations of traditional biochemical and computational methods. In this Roadmap, we highlight how artificial intelligence can transform our understanding of RNA biology by fostering collaborations between RNA biologists and computational scientists to drive innovation in this fundamental field of research. We discuss how non-coding regions of the mRNA, including introns and 5′ and 3′ untranslated regions, regulate the metabolism and interactomes of mRNA, and the current challenges in characterizing these regions. We further discuss large language models, which can be used to learn biologically meaningful RNA sequence representations. We also provide a detailed roadmap for integrating large language models with graph neural networks to harness publicly available sequencing and knowledge data. Adopting this roadmap will allow us to predict RNA interactions with diverse molecules and the modelling of context-specific mRNA interactomes.
-
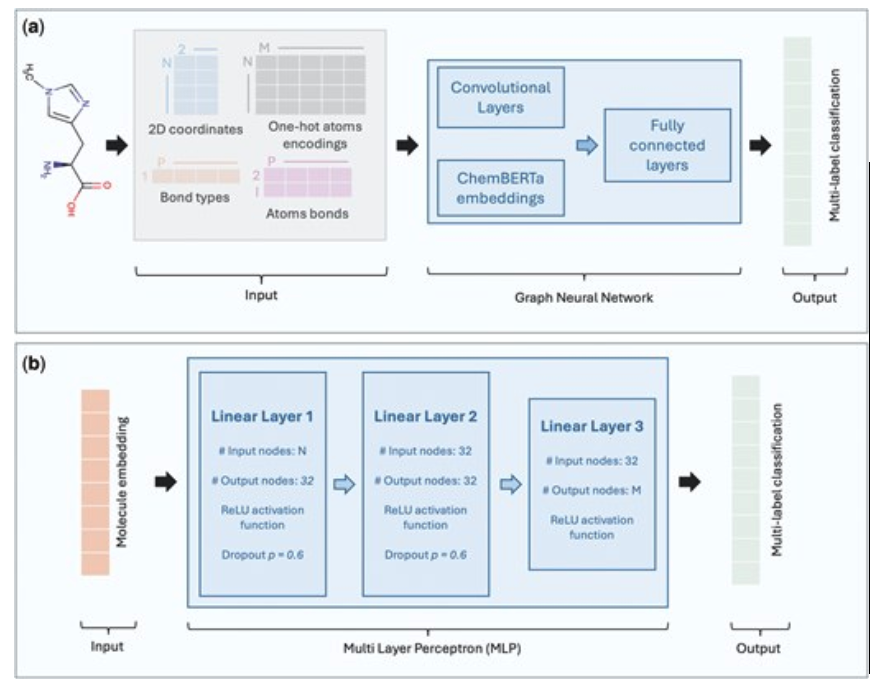 Structure-based metabolite function prediction using graph neural networksTancredi Cogne, Mariam Ait Oumelloul, Ali Saadat, Janna Hastings, and Jacques FellayBioinformatics Advances, Jul 2025
Structure-based metabolite function prediction using graph neural networksTancredi Cogne, Mariam Ait Oumelloul, Ali Saadat, Janna Hastings, and Jacques FellayBioinformatics Advances, Jul 2025Being able to broadly predict the function of novel metabolites based on their structures has applications in systems biology, environmental monitoring and drug discovery. To date, machine learning models aiming to predict functional characteristics of metabolites have largely been limited in scope to predicting single functions, or only a small number of functions simultaneously. Using the Human Metabolome Database as a source for a wider range of functional annotations, we assess the feasibility of predicting metabolite functions more broadly, as defined by four elements, namely location, role, the process it is involved in, and its physiological effect. We evaluated three graph neural network architectures to predict available functional ontology terms. We compared the graph models to two Multi-Layer Perceptron architectures using circular fingerprints and ChemBERTa embeddings. Among the models tested, the Graph Attention Network, incorporating embeddings from the pre-trained ChemBERTa model to predict the process metabolites are involved in, achieved the highest performance with a macro F1-score of 0.903 and an Area Under the Precision-Recall Curve of 0.926. The model identified function-associated structural patterns within metabolite families, demonstrating the potential for interpretable prediction of metabolite functions from structural information.
- A Bayesian Network Meta-analysis of Systemic Treatments for Metastatic Castration-Resistant Prostate Cancer in First- and Subsequent LinesMarie Wosny, Stefanie Aeppli, Stefanie Fischer, Tobias Peres, Christian Rothermundt, and Janna HastingsTargeted Oncology, Jun 2025
Metastatic castration-resistant prostate cancer (mCRPC) presents a challenge for clinicians in determining the optimal treatment sequence because of the lack of direct head-to-head comparisons, which is further complicated by the now-widespread use of androgen receptor pathway inhibitors (ARPIs) in metastatic hormone-sensitive prostate cancer (mHSPC).
-
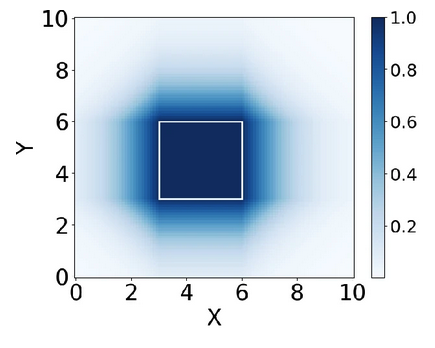 Box embeddings for extending ontologies: a data-driven and interpretable approachAdel Memariani, Martin Glauer, Simon Flügel, Fabian Neuhaus, Janna Hastings, and Till MossakowskiJournal of Cheminformatics, Sep 2025
Box embeddings for extending ontologies: a data-driven and interpretable approachAdel Memariani, Martin Glauer, Simon Flügel, Fabian Neuhaus, Janna Hastings, and Till MossakowskiJournal of Cheminformatics, Sep 2025Deriving symbolic knowledge from trained deep learning models is challenging due to the lack of transparency in such models. A promising approach to address this issue is to couple a semantic structure with the model outputs and thereby make the model interpretable. In prediction tasks such as multi-label classification, labels tend to form hierarchical relationships. Therefore, we propose enforcing a taxonomical structure on the model’s outputs throughout the training phase. In vector space, a taxonomy can be represented using axis-aligned hyper-rectangles, or boxes, which may overlap or nest within one another. The boundaries of a box determine the extent of a particular category. Thus, we used box-shaped embeddings of ontology classes to learn and transparently represent logical relationships that are only implicit in multi-label datasets. We assessed our model by measuring its ability to approximate the full set of inferred subclass relations in the ChEBI ontology, which is an important knowledge base in the field of life science. We demonstrate that our model captures implicit hierarchical relationships among labels, ensuring consistency with the underlying ontological conceptualization, while also achieving state-of-the-art performance in multi-label classification. Notably, this is accomplished without requiring an explicit taxonomy during the training process.
- Enhancing Interpretability of Ocular Disease Diagnosis: A Zero-Shot Study of Multimodal Large Language ModelsYating Pan and Janna HastingsStudies in Health Technology and Informatics, Aug 2025
Visual foundation models have advanced ocular disease diagnosis, yet providing interpretable explanations remains challenging. We evaluate multimodal LLMs for generating explanations of ocular diagnoses, combining Vision Transformer-derived saliency maps with clinical metadata. After finetuning RETFound for improved performance on the BRSET dataset (AUC-ROC 0.9664/0.8611 for diabetic retinopathy/glaucoma), we compared five LLMs through technical and clinical evaluations. GPT-o1 demonstrated superior performance across technical dimensions and clinical metrics (79.32% precision, 77.18% recall, 78.25% F1, 20.68% hallucination rate). Our findings highlight the importance of underlying diagnostic accuracy and advanced model architecture for generating reliable clinical explanations, suggesting opportunities for integrated verification mechanisms in future developments. The code and details can be found at: https://github.com/YatingPan/ocular-llm-explainability.
- Chemical classification program synthesis using generative artificial intelligenceChristopher J. Mungall, Adnan Malik, Daniel R. Korn, Justin T. Reese, Noel M. O’Boyle, and Janna HastingsJournal of Cheminformatics, Oct 2025
Accurately classifying chemical structures is essential for cheminformatics and bioinformatics, including tasks such as identifying bioactive compounds of interest, screening molecules for toxicity to humans, finding non-organic compounds with desirable material properties, or organizing large chemical libraries for drug discovery or environmental monitoring. However, manual classification is labor-intensive and difficult to scale to large chemical databases. Existing automated approaches either rely on manually constructed classification rules, or are deep learning methods that lack explainability. This work presents an approach that uses generative artificial intelligence to automatically write chemical classifier programs for classes in the Chemical Entities of Biological Interest (ChEBI) database. These programs can be used for efficient deterministic run-time classification of SMILES structures, with natural language explanations. The programs themselves constitute an explainable computable ontological model of chemical class nomenclature, which we call the ChEBI Chemical Class Program Ontology (C3PO). We validated our approach against the ChEBI database, and compared our results against deep learning models and a naive SMARTS pattern based classifier. C3PO outperforms the naive classifier, but does not reach the performance of state of the art deep learning methods. However, C3PO has a number of strengths that complement deep learning methods, including explainability and reduced data dependence. C3PO can be used alongside deep learning classifiers to provide an explanation of the classification, where both methods agree. The programs can be used as part of the ontology development process, and iteratively refined by expert human curators.
- Ontology pre-training improves machine learning-based predictions for metabolitesCharlotte Tumescheit, Martin Glauer, Simon Fluegel, Martin Larralde, Fabian Neuhaus, Till Mossakowski, and Janna HastingsOct 2025
Recent advances in the field of machine learning have shown that integration of expert knowledge improves performances, in particular for complex domains such as biology. Bio-ontologies offer a rich source of curated biological knowledge that can be harnessed to this end. Here, we describe an intuitive and generalisable approach to embed the knowledge contained in a classification hierarchy derived from a bio-ontology into a machine learning model as an intermediate training step between general-purpose pre-training and task-specific fine-tuning in a process that we call ’ontology pre-training’. We show that this approach leads to an improvement in predictive performance and a reduction in training time for a broad range of predictive tasks relevant to understanding metabolite functions in living systems, using a range of datasets derived from MoleculeNet. We see the biggest improvement for regression tasks, e.g. prediction of lipophilicity and aqueous solubility of molecules, and a robust improvement for most classification tasks. Our approach can be adapted for a wide range of knowledge sources, models and prediction tasks.
- Large Language Models Reveal Menstruation Experiences and Needs on Social MediaCharlotte Tumescheit, Davinny Sou, Marcia Nißen, Tobias Kowatsch, and Janna HastingsStudies in Health Technology and Informatics, Oct 2025
The gender knowledge gap in medicine, particularly regarding menstruation and disorders such as endometriosis, often results in delayed diagnoses and inadequate care. Many menstruating individuals report dismissal of debilitating symptoms, driving them to seek information and support on online platforms such as TikTok and YouTube. This study leverages social media to identify key topics reflecting lived experiences and needs to bridge this knowledge gap. Using a novel pipeline, we analysed video comments using BERTopic and the Llama 3.1 model. Key topics, including emotional support, educational guidance, and community validation, were consistent with prior research. This study underscores the potential of social media and large language models to inform inclusive menstrual health research, revealing unique insights regarding the menstruation experiences and needs of underrepresented and historically overlooked individuals such as those with irregular cycles.
- Application of a General Large Language Model-Based Classification System to Retrieve Information about Oncological TrialsFabio Dennstädt, Paul Windisch, Irina Filchenko, Johannes Zink, Paul Martin Putora, Ahmed Shaheen, Roberto Gaio, Nikola Cihoric, Marie Wosny, Stefanie Aeppli, and 3 more authorsOncology, Jun 2025
Introduction: The automated classification of clinical trials and key categories within the medical literature is increasingly relevant, particularly in oncology, as the volume of publications and trial reports continues to expand. Large language models (LLMs) may provide new opportunities for automating diverse classification tasks. They could be used for general-purpose text classification, retrieving information about oncological trials. Methods: A general text classification framework with adaptable prompt, model and categories for the classification was developed. The framework was tested with four datasets comprising nine binary classification questions related to oncological trials. Evaluation was conducted using a locally hosted Mixtral-8x7B-Instruct v0.1-GPTQ model and three cloud-based LLMs: Mixtral-8x7B-Instruct v0.1, Llama3.1-70B-Instruct, and Qwen-2.5–72B. Results: The system consistently produced valid responses with the local Mixtral-8x7B-Instruct model and the Llama3.1-70B-Instruct model. It achieved a response validity rate of 99.70% and 99.88% for the cloud-based Mixtral and Qwen models, respectively. Across all models, the framework achieved an overall accuracy of >94%, precision of >92%, recall of >90%, and an F1-score of >92%. Question-specific accuracy ranged from 86.33% to 99.83% for the local Mixtral model, 85.49%–99.83% for the cloud-based Mixtral model, 90.50%–99.83% for the Llama3.1 model, and 77.13%–99.83% for the Qwen model. Conclusion: The LLM-based classification framework exhibits robust accuracy and adaptability across various oncological trial classification tasks. While there remain some challenges such as strong prompt dependence and high computational and hardware demands, LLMs will play a crucial role in automating the classification of oncological trials and literature as the technology continues to advance.
- Deep learning aging marker from retinal images unveils sex-specific clinical and genetic signaturesOlga Trofimova, Leah Böttger, Sacha Bors, Yating Pan, Bart Liefers, Jose D. Vargas Quiros, Victor A. de Vries, Michael J. Beyeler, David M. Presby, Dennis Bontempi, and 3 more authorsOct 2025
Retinal fundus images offer a non-invasive window into systemic aging. Here, we fine-tuned a foundation model (RETFound) to predict chronological age from color fundus images in 71,343 participants from the UK Biobank, achieving a mean absolute error of 2.85 years. The resulting retinal age gap, i.e. the difference between predicted and chronological age, was associated with cardiometabolic traits, inflammation, cognitive performance, all-cause mortality, dementia, cancer, and incident cardiovascular disease. Genome-wide analyses identified genes related to longevity, metabolism, neurodegeneration, and age-related eye diseases. Sex-stratified models revealed consistent performance but divergent biological signatures: males had younger-appearing retinas and stronger links to metabolic syndrome, while in females, both model attention and genetic associations pointed to a greater involvement of retinal vasculature. Our study positions retinal aging as a biologically meaningful and sex-sensitive biomarker that can support more personalized approaches to risk assessment and aging-related healthcare.
- A comparative performance analysis of regular expressions and a large language model-based approach to extract the BI-RADS score from radiological reportsFabio Dennstädt, Luc Lerch, Max Schmerder, Nikola Cihoric, Grazia Maria Cereghetti, Roberto Gaio, Harald Bonel, Irina Filchenko, Janna Hastings, Florian Dammann, and 3 more authorsJAMIA Open, Dec 2025
Different natural language processing (NLP) techniques have demonstrated promising results for data extraction from radiological reports. Both traditional rule-based methods like regular expressions (Regex) and modern large language models (LLMs) can extract structured information. However, comparison between these approaches for extraction of specific radiological data elements has not been widely conducted.We compared accuracy and processing time between Regex and LLM-based approaches for extracting Breast Imaging-Reporting and Data System (BI-RADS) scores from 7764 radiology reports (mammography, ultrasound, MRI [magnetic resonance imaging], and biopsy). We developed a rule-based algorithm using Regex patterns and implemented an LLM-based extraction using the Rombos-LLM-V2.6-Qwen-14b model. A ground truth dataset of 199 manually classified reports was used for evaluation.We did not detect a statistically significant difference in the accuracy in extracting BI-RADS scores between Regex and an LLM-based method (accuracy of 89.20% for Regex vs 87.69% for the LLM-based method; P = .56, effect size w = 0.04; post-hoc power = 0.11). Compared to the LLM-based method, Regex processing was more efficient, completing the task 28 120 times faster (0.06 seconds vs 1687.20 seconds). Further analysis revealed that LLMs favored common classifications (particularly BI-RADS value of 2) while Regex more frequently returned “unclear” values. We also could confirm in our sample an already known laterality bias for breast cancer (BI-RADS 6) and detected a slight laterality skew for suspected breast cancer (BI-RADS 5) as well.For structured, standardized data like BI-RADS, traditional NLP techniques seem to be superior, though future work should explore hybrid approaches combining Regex precision for standardized elements with LLM contextual understanding for more complex information extraction tasks.This study compared 2 methods for automatically extracting information from medical reports. Thousands of radiology reports from breast cancer screenings were used to find the “Breast Imaging-Reporting and Data System (BI-RADS)” score, which helps doctors assess breast cancer risk. The goal was to see whether a traditional, rule-based method called Regular Expressions (Regex) or a newer artificial intelligence (AI) approach using large language models (LLMs) was better for finding the score in the reports. The scores of 199 reports were extracted by human researchers to find out how well the automated systems could to the same. The automated systems were then applied to the entire dataset with 7764 reports.The results showed that both methods were similarly accurate in finding the correct BI-RADS score. However, the simpler Regex method was much faster, completing the task over 28 000 times more quickly than the AI model.The study concludes that for extracting standardized data like the BI-RADS score, traditional techniques like Regex are more efficient and just as good as more complex AI. This suggests that the best approach for analyzing medical records may involve using a combination of older, faster tools for simple data and advanced AI for more complex information.
- Comparative Evaluation of a Medical Large Language Model in Answering Real-World Radiation Oncology Questions: Multicenter Observational StudyFabio Dennstädt, Max Schmerder, Elena Riggenbach, Lucas Mose, Katarina Bryjova, Nicolas Bachmann, Paul-Henry Mackeprang, Maiwand Ahmadsei, Dubravko Sinovcic, Paul Windisch, and 18 more authorsJournal of Medical Internet Research, Sep 2025
Background: Large language models (LLMs) hold promise for supporting clinical tasks, particularly in data-driven and technical disciplines such as radiation oncology. While prior evaluation studies have focused on examination-style settings for evaluating LLMs, their performance in real-life clinical scenarios remains unclear. In the future, LLMs might be used as general AI assistants to answer questions arising in clinical practice. It is unclear how well a modern LLM, locally executed within the infrastructure of a hospital, would answer such questions compared with clinical experts. Objective: This study aimed to assess the performance of a locally deployed, state-of-the-art medical LLM in answering real-world clinical questions in radiation oncology compared with clinical experts. The aim was to evaluate the overall quality of answers, as well as the potential harmfulness of the answers if used for clinical decision-making. Methods: Physicians from 10 departments of European hospitals collected questions arising in the clinical practice of radiation oncology. Fifty of these questions were answered by 3 senior radiation oncology experts with at least 10 years of work experience, as well as the LLM Llama3-OpenBioLLM-70B (Ankit Pal and Malaikannan Sankarasubbu). In a blinded review, physicians rated the overall answer quality on a 5-point Likert scale (quality), assessed whether an answer might be potentially harmful if used for clinical decision-making (harmfulness), and determined if responses were from an expert or the LLM (recognizability). Comparisons between clinical experts and LLMs were then made for quality, harmfulness, and recognizability. Results: There were no significant differences between the quality of the answers between LLM and clinical experts (mean scores of 3.38 vs 3.63; median 4.00, IQR 3.00-4.00 vs median 3.67, IQR 3.33-4.00; P=.26; Wilcoxon signed rank test). The answers were deemed potentially harmful in 13% of cases for the clinical experts compared with 16% of cases for the LLM (P=.63; Fisher exact test). Physicians correctly identified whether an answer was given by a clinical expert or an LLM in 78% and 72% of cases, respectively. Conclusions: A state-of-the-art medical LLM can answer real-life questions from the clinical practice of radiation oncology similarly well as clinical experts regarding overall quality and potential harmfulness. Such LLMs can already be deployed within the local hospital environment at an affordable cost. While LLMs may not yet be ready for clinical implementation as general AI assistants, the technology continues to improve at a rapid pace. Evaluation studies based on real-life situations are important to better understand the weaknesses and limitations of LLMs in clinical practice. Such studies are also crucial to define when the technology is ready for clinical implementation. Furthermore, education for health care professionals on generative AI is needed to ensure responsible clinical implementation of this transforming technology.
2024
- Preventing harm from non-conscious bias in medical generative AIJanna HastingsThe Lancet Digital Health, Jan 2024
- Interpretable ontology extension in chemistryMartin Glauer, Adel Memariani, Fabian Neuhaus, Till Mossakowski, and Janna HastingsSemantic Web, Jan 2024
Reference ontologies provide a shared vocabulary and knowledge resource for their domain. Manual construction and annotation enables them to maintain high quality, allowing them to be widely accepted across their community. However, the manual ontolo
- Chebifier: automating semantic classification in ChEBI to accelerate data-driven discoveryMartin Glauer, Fabian Neuhaus, Simon Flügel, Marie Wosny, Till Mossakowski, Adel Memariani, Johannes Schwerdt, and Janna HastingsDigital Discovery, Jan 2024
- Cross-Care: Assessing the Healthcare Implications of Pre-training Data on Language Model BiasShan Chen, Jack Gallifant, Mingye Gao, Pedro Moreira, Nikolaj Munch, Ajay Muthukkumar, Arvind Rajan, Jaya Kolluri, Amelia Fiske, Janna Hastings, and 5 more authorsIn , May 2024
Large language models (LLMs) are increasingly essential in processing natural languages, yet their application is frequently compromised by biases and inaccuracies originating in their training data. In this study, we introduce Cross-Care, the first benchmark framework dedicated to assessing biases and real world knowledge in LLMs, specifically focusing on the representation of disease prevalence across diverse demographic groups. We systematically evaluate how demographic biases embedded in pre-training corpora like \ThePile influence the outputs of LLMs. We expose and quantify discrepancies by juxtaposing these biases against actual disease prevalences in various U.S. demographic groups. Our results highlight substantial misalignment between LLM representation of disease prevalence and real disease prevalence rates across demographic subgroups, indicating a pronounced risk of bias propagation and a lack of real-world grounding for medical applications of LLMs. Furthermore, we observe that various alignment methods minimally resolve inconsistencies in the models’ representation of disease prevalence across different languages. For further exploration and analysis, we make all data and a data visualization tool available at: www.crosscare.net.
-
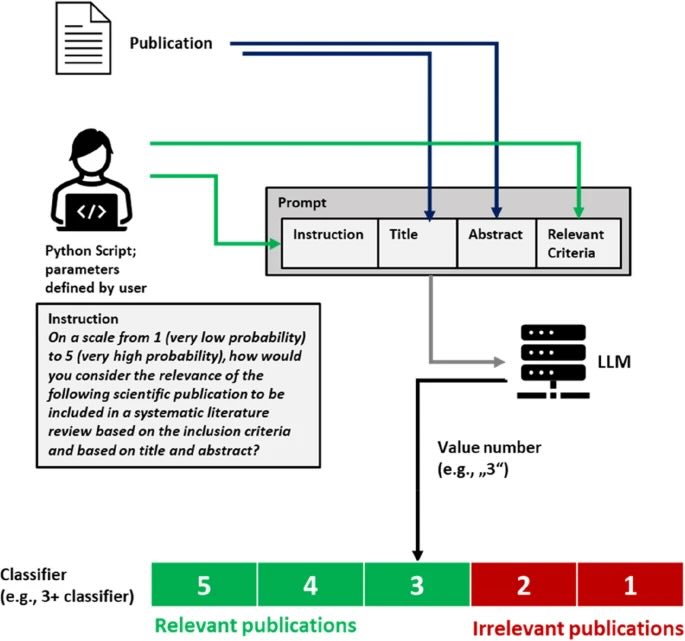 Title and abstract screening for literature reviews using large language models: an exploratory study in the biomedical domainFabio Dennstädt, Johannes Zink, Paul Martin Putora, Janna Hastings, and Nikola CihoricSystematic Reviews, Jun 2024
Title and abstract screening for literature reviews using large language models: an exploratory study in the biomedical domainFabio Dennstädt, Johannes Zink, Paul Martin Putora, Janna Hastings, and Nikola CihoricSystematic Reviews, Jun 2024Systematically screening published literature to determine the relevant publications to synthesize in a review is a time-consuming and difficult task. Large language models (LLMs) are an emerging technology with promising capabilities for the automation of language-related tasks that may be useful for such a purpose.
-
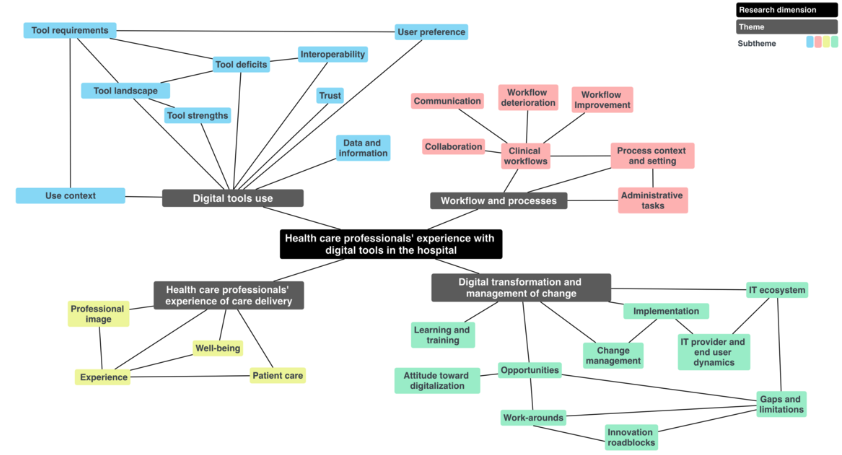 The Paradoxes of Digital Tools in Hospitals: Qualitative Interview StudyMarie Wosny, Livia Maria Strasser, and Janna HastingsJournal of Medical Internet Research, Jul 2024
The Paradoxes of Digital Tools in Hospitals: Qualitative Interview StudyMarie Wosny, Livia Maria Strasser, and Janna HastingsJournal of Medical Internet Research, Jul 2024Background: Digital tools are progressively reshaping the daily work of health care professionals (HCPs) in hospitals. While this transformation holds substantial promise, it leads to frustrating experiences, raising concerns about negative impacts on clinicians’ well-being. Objective: The goal of this study was to comprehensively explore the lived experiences of HCPs navigating digital tools throughout their daily routines. Methods: Qualitative in-depth interviews with 52 HCPs representing 24 medical specialties across 14 hospitals in Switzerland were performed. Results: Inductive thematic analysis revealed 4 main themes: digital tool use, workflow and processes, HCPs’ experience of care delivery, and digital transformation and management of change. Within these themes, 6 intriguing paradoxes emerged, and we hypothesized that these paradoxes might partly explain the persistence of the challenges facing hospital digitalization: the promise of efficiency and the reality of inefficiency, the shift from face to face to interface, juggling frustration and dedication, the illusion of information access and trust, the complexity and intersection of workflows and care paths, and the opportunities and challenges of shadow IT. Conclusions: Our study highlights the central importance of acknowledging and considering the experiences of HCPs to support the transformation of health care technology and to avoid or mitigate any potential negative experiences that might arise from digitalization. The viewpoints of HCPs add relevant insights into long-standing informatics problems in health care and may suggest new strategies to follow when tackling future challenges.
- Factors Guiding Clinical Decision-Making in Genitourinary OncologyMarie Wosny, Stefanie Aeppli, Stefanie Fischer, Tobias Peres, Christian Rothermundt, and Janna HastingsCancer Medicine, Jul 2024
Introduction Clinical decision-making in oncology is a complex process, with the primary goal of identifying the most effective treatment tailored to individual cancer patients. Many factors influence the treatment decision: disease- and patient-specific criteria, the increasingly complex treatment landscape, market authorization and drug availability, financial aspects, and personal treatment expertise. In the domain of genitourinary cancers, particularly prostate cancer, decision-making is challenging. Despite the prevalence of this malignancy, there are few in-depth explorations of these factors within real-world scenarios. Understanding and refining this intricate decision-making process is essential for future successful clinical decisions and the integration of computerized decision support into clinicians’ workflows. Aim The objective of this study is to improve the current knowledge base and evidence of the factors that influence treatment decision-making for patients with genitourinary cancers. Methods Assessment of how routine treatment decisions are made for genitourinary cancers was performed by a mixed-methods study, encompassing field observations and focus group discussions. Results In total, we identified 59 factors that influence clinical decision-making in oncology, specifically for genitourinary and prostate cancer. Of these, 23 criteria can be classified as decision-maker-related criteria encompassing personal, cognitive, and emotional attributes and factors of both, healthcare professionals and patients. Moreover, 20 decision-specific criteria have been identified that refer to clinical and disease-related factors, followed by 16 contextual decision factors that describe the relevant criteria introduced by the specific circumstances and environment in which the treatment decision is made. Conclusion By presenting an exhaustive set of decision factors and providing specific examples for genitourinary cancers, this observational study establishes a possible framework for a better understanding of decision-making. Moreover, we specify and expand the set of decision factors, while emphasizing the importance of cognitive, emotional, and human factors, as well as the quality and accessibility of decision-relevant information.
- Evaluating Text-to-Image Generated Photorealistic Images of Human AnatomyPaula Muhr, Yating Pan, Charlotte Tumescheit, Ann-Kathrin Kübler, Hatice Kübra Parmaksiz, Cheng Chen, Pablo Sebastián Bolaños Orozco, Soeren S. Lienkamp, Janna Hastings, Paula Muhr, and 8 more authorsCureus, Nov 2024
Background: Generative artificial intelligence (AI) models that can produce photorealistic images from text descriptions have many applications in medicine, including medical education and the generation of synthetic data. However, it can be challenging to evaluate their heterogeneous outputs and to compare between different models. There is a need for a systematic approach enabling image and model comparisons. Method: To address this gap, we developed an error classification system for annotating errors in AI-generated photorealistic images of humans and applied our method to a corpus of 240 images generated with three different models (DALL-E 3, Stable Diffusion XL, and Stable Cascade) using 10 prompts with eight images per prompt. Results: The error classification system identifies five different error types with three different severities across five anatomical regions and specifies an associated quantitative scoring method based on aggregated proportions of errors per expected count of anatomical components for the generated image. We assessed inter-rater agreement by double-annotating 25% of the images and calculating Krippendorf’s alpha and compared results across the three models and 10 prompts quantitatively using a cumulative score per image. The error classification system, accompanying training manual, generated image collection, annotations, and all associated scripts, is available from our GitHub repository at https://github.com/hastingslab-org/ai-human-images. Inter-rater agreement was relatively poor, reflecting the subjectivity of the error classification task. Model comparisons revealed that DALL-E 3 performed consistently better than Stable Diffusion; however, the latter generated images reflecting more diversity in personal attributes. Images with groups of people were more challenging for all the models than individuals or pairs; some prompts were challenging for all models. Conclusion: Our method enables systematic comparison of AI-generated photorealistic images of humans; our results can serve to catalyse improvements in these models for medical applications.
- Electronic cigarettes and subsequent use of cigarettes in young people: An evidence and gap mapMonserrat Conde, Kate Tudor, Rachna Begh, Rebecca Nolan, Sufen Zhu, Dimitra Kale, Sarah Jackson, Jonathan Livingstone-Banks, Nicola Lindson, Caitlin Notley, and 5 more authorsAddiction, Nov 2024
Background and aims The use of e-cigarettes may influence later smoking uptake in young people. Evidence and gap maps (EGMs) are interactive on-line tools that display the evidence and gaps in a specific area of policy or research. The aim of this study was to map clusters and gaps in evidence exploring the relationship between e-cigarette use or availability and subsequent combustible tobacco use in people aged \textless 30 years. Methods We conducted an EGM of primary studies and systematic reviews. A framework and an interactive EGM was developed in consultation with an expert advisory group. A systematic search of five databases retrieved 9057 records, from which 134 studies were included. Systematic reviews were appraised using AMSTAR-2, and all included studies were coded into the EGM framework resulting in the interactive web-based EGM. A descriptive analysis of key characteristics of the identified evidence clusters and gaps resulted in this report. Results Studies were completed between 2015 and 2023, with the first systematic reviews being published in 2017. Most studies were conducted in western high-income countries, predominantly the United States. Cohort studies were the most frequently used study design. The evidence is clustered on e-cigarette use as an exposure, with an absolute gap identified for evidence looking into the availability of e-cigarettes and subsequent cessation of cigarette smoking. We also found little evidence analysing equity factors, and little exploring characteristics of e-cigarette devices. Conclusions This evidence and gap map (EGM) offers a tool to explore the available evidence regarding the e-cigarette use/availability and later cigarette smoking in people under the age of 30 years at the time of the search. The majority of the 134 reports is from high-income countries, with an uneven geographic distribution. Most of the systematic reviews are of lower quality, suggesting the need for higher-quality reviews. The evidence is clustered around e-cigarette use as an exposure and subsequent frequency/intensity of current combustible tobacco use. Gaps in evidence focusing on e-cigarette availability, as well as on the influence of equity factors may warrant further research. This EGM can support funders and researchers in identifying future research priorities, while guiding practitioners and policymakers to the current evidence base.
- Proteomic profiling for biomarker discovery in heparin-induced thrombocytopeniaHenning Nilius, Hind Hamzeh-Cognasse, Janna Hastings, Jan-Dirk Studt, Dimitrios A. Tsakiris, Andreas Greinacher, Adriana Mendez, Adrian Schmidt, Walter A. Wuillemin, Bernhard Gerber, and 6 more authorsBlood Advances, May 2024
New analytical techniques can assess hundreds of proteins simultaneously with high sensitivity, facilitating the observation of their complex interplay and role in disease mechanisms. We hypothesized that proteomic profiling targeting proteins involved in thrombus formation, inflammation, and the immune response would identify potentially new biomarkers for heparin-induced thrombocytopenia (HIT). Four existing panels of the Olink proximity extension assay covering 356 proteins involved in thrombus formation, inflammation, and immune response were applied to randomly selected patients with suspected HIT (confirmed HIT, n = 32; HIT ruled out, n = 38; and positive heparin/platelet factor 4 [H/PF4] antibodies, n = 28). The relative difference in protein concentration was analyzed using a linear regression model adjusted for sex and age. To confirm the test results, soluble P-selectin was determined using enzyme-linked immunosorbent assay (ELISA) in above mentioned patients and an additional second data set (n = 49). HIT was defined as a positive heparin-induced platelet activation assay (washed platelet assay). Among 98 patients of the primary data set, the median 4Ts score was 5 in patients with HIT, 4 in patients with positive H/PF4 antibodies, and 3 in patients without HIT. The median optical density of a polyspecific H/PF4 ELISA were 3.0, 0.9, and 0.3. Soluble P-selectin remained statistically significant after multiple test adjustments. The area under the receiver operating characteristic curve was 0.81 for Olink and 0.8 for ELISA. Future studies shall assess the diagnostic and prognostic value of soluble P-selectin in the management of HIT.
- Practical Recommendations for Navigating Digital Tools in Hospitals: Qualitative Interview StudyMarie Wosny, Livia Maria Strasser, Simone Kraehenmann, and Janna HastingsJMIR Medical Education, Nov 2024
Background: The digitalization of health care organizations is an integral part of a clinician’s daily life, making it vital for health care professionals (HCPs) to understand and effectively use digital tools in hospital settings. However, clinicians often express a lack of preparedness for their digital work environments. Particularly, new clinical end users, encompassing medical and nursing students, seasoned professionals transitioning to new health care environments, and experienced practitioners encountering new health care technologies, face critically intense learning periods, often with a lack of adequate time for learning digital tools, resulting in difficulties in integrating and adopting these digital tools into clinical practice. Objective: This study aims to comprehensively collect advice from experienced HCPs in Switzerland to guide new clinical end users on how to initiate their engagement with health ITs within hospital settings. Methods: We conducted qualitative interviews with 52 HCPs across Switzerland, representing 24 medical specialties from 14 hospitals. The interviews were transcribed verbatim and analyzed through inductive thematic analysis. Codes were developed iteratively, and themes and aggregated dimensions were refined through collaborative discussions. Results: Ten themes emerged from the interview data, namely (1) digital tool understanding, (2) peer-based learning strategies, (3) experimental learning approaches, (4) knowledge exchange and support, (5) training approaches, (6) proactive innovation, (7) an adaptive technology mindset, (8) critical thinking approaches, (9) dealing with emotions, and (10) empathy and human factors. Consequently, we devised 10 recommendations with specific advice to new clinical end users on how to approach new health care technologies, encompassing the following: take time to get to know and understand the tools you are working with; proactively ask experienced colleagues; simply try it out and practice; know where to get help and information; take sufficient training; embrace curiosity and pursue innovation; maintain an open and adaptable mindset; keep thinking critically and use your knowledge base; overcome your fears, and never lose the human and patient focus. Conclusions: Our study emphasized the importance of comprehensive training and learning approaches for health care technologies based on the advice and recommendations of experienced HCPs based in Swiss hospitals. Moreover, these recommendations have implications for medical educators and clinical instructors, providing advice on effective methods to instruct and support new end users, enabling them to use novel technologies proficiently. Therefore, we advocate for new clinical end users, health care institutions and clinical instructors, academic institutions and medical educators, and regulatory bodies to prioritize effective training and cultivating technological readiness to optimize IT use in health care.
- Application of a general LLM-based classification system to retrieve information about oncological trialsFabio Dennstädt, Paul Windisch, Irina Filchenko, Johannes Zink, Paul Martin Putora, Ahmed Shaheen, Roberto Gaio, Nikola Cihoric, Marie Wosny, Stefanie Aeppli, and 3 more authorsDec 2024
Purpose The automated classification of clinical trials and medical literature is increasingly relevant, particularly in oncology, as the volume of publications and trial reports continues to expand. Large Language Models (LLMs) may provide new opportunities for automated diverse classification tasks. In this study, we developed a general-purpose text classification framework using LLMs and evaluated its performance on oncological trial classification tasks. Methods and Materials A general text classification framework with adaptable prompt, model and categories for the classification was developed. The framework was tested with four datasets comprising nine binary classification questions related to oncological trials. Evaluation was conducted using a locally hosted version of Mixtral-8x7B-Instruct v0.1 and three cloud-based LLMs: Mixtral-8x7B-Instruct v0.1, Llama3.1-70B-Instruct, and Qwen-2.5-72B. Results The system consistently produced valid responses with the local Mixtral-8x7B-Instruct model and the Llama3.1-70B-Instruct model. It achieved a response validity rate of 99.70% and 99.88% for the cloud-based Mixtral and Qwen models, respectively. Across all models, the framework achieved an overall accuracy of \textgreater94%, precision of \textgreater92%, recall of \textgreater90%, and an F1-score of \textgreater92%. Question-specific accuracy ranged from 86.33% to 99.83% for the local Mixtral model, 85.49% to 99.83% for the cloud-based Mixtral model, 90.50% to 99.83% for the Llama3.1 model, and 77.13% to 99.83% for the Qwen model. Conclusions The LLM-based classification framework exhibits robust accuracy and adaptability across various oncological trial classification tasks. The findings highlight the potential of automated, LLM- driven trial classification systems, which may become increasingly used in oncology.
- Horned-OWL: Flying Further and Faster with OntologiesPhillip Lord, Björn Gehrke, Martin Larralde, Janna Hastings, Filippo De Bortoli, James A. Overton, James P. Balhoff, and Jennifer WarrenderTransactions on Graph Data and Knowledge, Dec 2024
- Trace amine-associated receptor 1 (TAAR1) agonism for psychosis: a living systematic review and meta-analysis of human and non-human dataSpyridon Siafis, Virginia Chiocchia, Malcolm R. Macleod, Charlotte Austin, Ava Homiar, Francesca Tinsdeall, Claire Friedrich, Fiona J. Ramage, Jaycee Kennett, Nobuyuki Nomura, and 29 more authorsWellcome Open Research, Dec 2024
BACKGROUND: Trace amine-associated receptor 1 (TAAR1) agonism shows promise for treating psychosis, prompting us to synthesise data from human and non-human studies. METHODS: We co-produced a living systematic review of controlled studies examining TAAR1 agonists in individuals (with or without psychosis/schizophrenia) and relevant animal models. Two independent reviewers identified studies in multiple electronic databases (until 17.11.2023), extracted data, and assessed risk of bias. Primary outcomes were standardised mean differences (SMD) for overall symptoms in human studies and hyperlocomotion in animal models. We also examined adverse events and neurotransmitter signalling. We synthesised data with random-effects meta-analyses. RESULTS: Nine randomised trials provided data for two TAAR1 agonists (ulotaront and ralmitaront), and 15 animal studies for 10 TAAR1 agonists. Ulotaront and ralmitaront demonstrated few differences compared to placebo in improving overall symptoms in adults with acute schizophrenia (N=4 studies, n=1291 participants; SMD=0.15, 95%CI: -0.05, 0.34), and ralmitaront was less efficacious than risperidone (N=1, n=156, SMD=-0.53, 95%CI: -0.86, -0.20). Large placebo response was observed in ulotaront phase-III trials. Limited evidence suggested a relatively benign side-effect profile for TAAR1 agonists, although nausea and sedation were common after a single dose of ulotaront. In animal studies, TAAR1 agonists improved hyperlocomotion compared to control (N=13 studies, k=41 experiments, SMD=1.01, 95%CI: 0.74, 1.27), but seemed less efficacious compared to dopamine D 2 receptor antagonists (N=4, k=7, SMD=-0.62, 95%CI: -1.32, 0.08). Limited human and animal data indicated that TAAR1 agonists may regulate presynaptic dopaminergic signalling. CONCLUSIONS: TAAR1 agonists may be less efficacious than dopamine D 2 receptor antagonists already licensed for schizophrenia. The results are preliminary due to the limited number of drugs examined, lack of longer-term data, publication bias, and assay sensitivity concerns in trials associated with large placebo response. Considering their unique mechanism of action, relatively benign side-effect profile and ongoing drug development, further research is warranted. REGISTRATION: PROSPERO-ID: CRD42023451628.
- Towards an ontology of mental health: Protocol for developing an ontology to structure and integrate evidence regarding anxiety, depression and psychosisPaulina M. Schenk, Janna Hastings, Micaela Santilli, Jennifer Potts, Jaycee Kennett, Claire Friedrich, and Susan MichieWellcome Open Research, Dec 2024
BACKGROUND: Research about anxiety, depression and psychosis and their treatments is often reported using inconsistent language, and different aspects of the overall research may be conducted in separate silos. This leads to challenges in evidence synthesis and slows down the development of more effective interventions to prevent and treat these conditions. To address these challenges, the Global Alliance for Living Evidence on aNxiety, depressiOn and pSychosis (GALENOS) Project is conducting a series of living systematic reviews about anxiety, depression and psychosis. An ontology (a classification and specification framework) for the domain of mental health is being created to organise and synthesise evidence within these reviews and present them in a structured online data repository. AIM: This study aims to develop an ontology of mental health that includes entities with clear labels and definitions to describe and synthesise evidence about mental health, focusing on anxiety, depression and psychosis. METHODS: We will develop and apply the GALENOS Mental Health Ontology through eight steps: (1) defining the ontology’s scope; (2) identifying, labelling and defining the ontology’s entities for the GALENOS living systematic reviews; (3) structuring the ontology’s upper level (4) refining the upper level’s clarity and scope via a stakeholder consultation; (5) formally specifying the relationships between entities in the Mental Health Ontology; (6) making the ontology machine-readable and available online; (7) integrating the ontology into the data repository; and (8) exploring the ontology-structured repository’s usability. CONCLUSION AND DISCUSSION: The Mental Health Ontology supports the formal representation of complex upper-level entities within mental health and their relationships. It will enable more explicit and precise communication and evidence synthesis about anxiety, depression and psychosis across the GALENOS Project’s living systematic reviews. By being computer readable, the ontology can also be harnessed within algorithms that support automated categorising, linking, retrieving and synthesising evidence.
- Applying Large Language Models to Interpret Qualitative Interviews in HealthcareMarie Wosny and Janna HastingsStudies in Health Technology and Informatics, Aug 2024
To address the persistent challenges in healthcare, it is crucial to incorporate firsthand experiences and perspectives from stakeholders such as patients and healthcare professionals. However, the current process of collecting, analyzing and interpreting qualitative data, such as interviews, is slow and labor-intensive. To expedite this process and enhance efficiency, automated approaches aim to extract meaningful themes and accelerate interpretation, but current approaches such as topic modeling reduce the richness of the raw data. Here, we evaluate whether Large Language Models can be used to support the semi-automated interpretation of qualitative interview data. We compare a novel approach based on LLMs to topic modeling approaches and to manually identified themes across two different qualitative interview datasets. This exploratory study finds that LLMs have the potential to support incorporating human perspectives more widely in the advancement of sustainable healthcare systems.
2023
- Ontology Pre-training for Poison PredictionMartin Glauer, Fabian Neuhaus, Till Mossakowski, and Janna HastingsIn Proc. KI 2023, Jan 2023
Integrating human knowledge into neural networks has the potential to improve their robustness and interpretability. We have developed a novel approach to integrate knowledge from ontologies into the structure of a Transformer network which we call ontology pre-training: we train the network to predict membership in ontology classes as a way to embed the structure of the ontology into the network, and subsequently fine-tune the network for the particular prediction task. We apply this approach to a case study in predicting the potential toxicity of a small molecule based on its molecular structure, a challenging task for machine learning in life sciences chemistry. Our approach improves on the state of the art, and moreover has several additional benefits. First, we are able to show that the model learns to focus attention on more meaningful chemical groups when making predictions with ontology pre-training than without, paving a path towards greater robustness and interpretability. Second, the training time is reduced after ontology pre-training, indicating that the model is better placed to learn what matters for toxicity prediction with the ontology pre-training than without. This strategy has general applicability as a neuro-symbolic approach to embed meaningful semantics into neural networks.
- Using machine learning to extract information and predict outcomes from reports of randomised trials of smoking cessation interventions in the Human Behaviour Change ProjectRobert West, Francesca Bonin, James Thomas, Alison J. Wright, Pol Mac Aonghusa, Martin Gleize, Yufang Hou, Alison O’Mara-Eves, Janna Hastings, Marie Johnston, and 1 more authorWellcome Open Research, Jan 2023
- An ontology of mechanisms of action in behaviour change interventionsPaulina Schenk, Alison Wright, Robert West, Janna Hastings, Fabiana Lorencatto, Candice Moore, Emily Hayes, Verena Schneider, Ella Howes, and Susan MichieApr 2023
Background: Behaviour change interventions influence behaviour through causal processes called “mechanisms of action” (MoAs). Reports of such interventions and their evaluations often use inconsistent or ambiguous terminology. This includes the reporting of MoAs, creating challenges for searching, evidence synthesis and theory development. An ontology can help address these challenges by serving as a classification system that labels and defines classes for MoAs and their relationships. Aim: To develop an ontology of MoAs of behaviour change interventions. Methods: To develop the MoA Ontology, we (1) defined the ontology’s scope; (2) identified, labelled and defined the ontology’s classes; (3) refined the ontology by annotating (i.e., coding) MoAs in intervention reports; (4) refined the ontology via stakeholder review of the ontology’s comprehensiveness and clarity; (5) tested whether researchers could reliably apply the ontology to annotate MoAs in intervention reports; (6) refined the relationships between classes; (7) reviewed the alignment of the MoA Ontology with relevant ontologies, (8) reviewed the ontology’s alignment with the Theories and Techniques Project; and (9) published the ontology and created a machine-readable version. Results: An MoA was defined as “a process that is causally active in the relationship between a behaviour change intervention scenario and its outcome behaviour”. We created an initial MoA Ontology with 261 classes through Steps 2-5. Inter-rater reliability for annotating study reports using these classes was α=0.68 (“acceptable”) for researchers familiar with the ontology and α=0.47 for researchers unfamiliar with it. As a result of additional revisions (Steps 6-8), 21 further classes were added to the ontology resulting in 282 classes organised into seven hierarchical levels. Conclusions and implications: The MoA Ontology extensively captures MoAs of behaviour change interventions. The ontology can serve as a controlled vocabulary for MoAs to consistently describe and synthesise evidence about MoAs across diverse sources.
- New living evidence resource of human and non-human studies for early intervention and research prioritisation in anxiety, depression and psychosisAndrea Cipriani, Soraya Seedat, Lea Milligan, Georgia Salanti, Malcolm Macleod, Janna Hastings, James Thomas, Susan Michie, Toshi A. Furukawa, David Gilbert, and 24 more authorsBMJ Ment Health, Jun 2023
In anxiety, depression and psychosis, there has been frustratingly slow progress in developing novel therapies that make a substantial difference in practice, as well as in predicting which treatments will work for whom and in what contexts. To intervene early in the process and deliver optimal care to patients, we need to understand the underlying mechanisms of mental health conditions, develop safe and effective interventions that target these mechanisms, and improve our capabilities in timely diagnosis and reliable prediction of symptom trajectories. Better synthesis of existing evidence is one way to reduce waste and improve efficiency in research towards these ends. Living systematic reviews produce rigorous, up-to-date and informative evidence summaries that are particularly important where research is emerging rapidly, current evidence is uncertain and new findings might change policy or practice. Global Alliance for Living Evidence on aNxiety, depressiOn and pSychosis (GALENOS) aims to tackle the challenges of mental health science research by cataloguing and evaluating the full spectrum of relevant scientific research including both human and preclinical studies. GALENOS will also allow the mental health community—including patients, carers, clinicians, researchers and funders—to better identify the research questions that most urgently need to be answered. By creating open-access datasets and outputs in a state-of-the-art online resource, GALENOS will help identify promising signals early in the research process. This will accelerate translation from discovery science into effective new interventions for anxiety, depression and psychosis, ready to be translated in clinical practice across the world.
- Human Factors Influencing the Experience of Healthcare Professionals Using Digital ToolsMarie Wosny, Livia Strasser, and Janna HastingsStudies in health technology and informatics, Jun 2023
Digitalization in healthcare has the potential to offer numerous advantages to various stakeholders, however, healthcare professionals often encounter difficulties while using digital tools. We conducted a qualitative analysis of published studies to examine the experience of clinicians using digital tools. Our findings revealed that human factors influence clinicians’ experiences and that integration of human factors into the design and development of healthcare technologies is of high importance to improve user experience and overall success.
- Experience of Health Care Professionals Using Digital Tools in the Hospital: Qualitative Systematic ReviewMarie Wosny, Livia Maria Strasser, and Janna HastingsJMIR Human Factors, Oct 2023
Background: The digitalization of health care has many potential benefits, but it may also negatively impact health care professionals’ well-being. Burnout can, in part, result from inefficient work processes related to the suboptimal implementation and use of health information technologies. Although strategies to reduce stress and mitigate clinician burnout typically involve individual-based interventions, emerging evidence suggests that improving the experience of using health information technologies can have a notable impact. Objective: The aim of this systematic review was to collect evidence of the benefits and challenges associated with the use of digital tools in hospital settings with a particular focus on the experiences of health care professionals using these tools. Methods: We conducted a systematic literature review following the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines to explore the experience of health care professionals with digital tools in hospital settings. Using a rigorous selection process to ensure the methodological quality and validity of the study results, we included qualitative studies with distinct data that described the experiences of physicians and nurses. A panel of 3 independent researchers performed iterative data analysis and identified thematic constructs. Results: Of the 1175 unique primary studies, we identified 17 (1.45%) publications that focused on health care professionals’ experiences with various digital tools in their day-to-day practice. Of the 17 studies, 10 (59%) focused on clinical decision support tools, followed by 6 (35%) studies focusing on electronic health records and 1 (6%) on a remote patient-monitoring tool. We propose a theoretical framework for understanding the complex interplay between the use of digital tools, experience, and outcomes. We identified 6 constructs that encompass the positive and negative experiences of health care professionals when using digital tools, along with moderators and outcomes. Positive experiences included feeling confident, responsible, and satisfied, whereas negative experiences included frustration, feeling overwhelmed, and feeling frightened. Positive moderators that may reinforce the use of digital tools included sufficient training and adequate workflow integration, whereas negative moderators comprised unfavorable social structures and the lack of training. Positive outcomes included improved patient care and increased workflow efficiency, whereas negative outcomes included increased workload, increased safety risks, and issues with information quality. Conclusions: Although positive and negative outcomes and moderators that may affect the use of digital tools were commonly reported, the experiences of health care professionals, such as their thoughts and emotions, were less frequently discussed. On the basis of this finding, this study highlights the need for further research specifically targeting experiences as an important mediator of clinician well-being. It also emphasizes the importance of considering differences in the nature of specific tools as well as the profession and role of individual users. Trial Registration: PROSPERO CRD42023393883; https://tinyurl.com/2htpzzxj
- Exploring the capabilities of Large Language Models such as ChatGPT in radiation oncologyFabio Dennstädt, Janna Hastings, Paul Martin Putora, Erwin Vu, Galina Fischer, Krisztian Süveg, Markus Glatzer, Elena Riggenbach, Hông-Linh Hà, and Nikola CihoricAdvances in Radiation Oncology, Nov 2023
Purpose Technological progress of machine learning and natural language processing (NLP) led to the development of large language models (LLMs), capable of producing well-formed text responses and providing natural language access to knowledge. Modern conversational LLMs such as ChatGPT have shown remarkable capabilities across a variety of fields, including medicine. These models may assess even highly specialized medical knowledge within specific disciplines, such as radiation therapy. We conducted an exploratory study to examine the capabilities of ChatGPT to answer questions in radiation therapy. Methods and Materials A set of multiple-choice questions about clinical, physics and biology general knowledge in radiation oncology as well as a set of open-ended questions were created. These were given as prompts to the LLM ChatGPT, and the answers were collected and analyzed. For the multiple-choice questions, it was checked how many of the answers could be clearly assigned to one of the answers and the portion of correct answers was determined. For the open-ended questions, independent blinded radiation oncologists evaluated the quality of the answers regarding correctness and usefulness on a 5-point Likert scale. Furthermore, the evaluators were asked to provide suggestions for improving the quality of the answers. Results For 70 multiple-choice questions, ChatGPT gave valid answers in 66 cases (94.3%). In 60.61% of the valid answers, the selected answer was correct (50.0% of clinical questions, 78.6% of physics questions and 58.3% of biology questions). For 25 open-ended questions, 12 answers of ChatGPT were considered as “acceptable”, “good” or “very good” regarding both correctness and helpfulness by all six participating radiation oncologists. Overall, the answers were considered “very good” in 29.3%/28%, “good” in 28%/29.3%, “acceptable” in 19.3%/19.3%, “bad” in 9.3%/9.3% and “very bad” in 14%/14% regarding correctness/helpfulness. Conclusions Modern conversational LLMs such as ChatGPT can provide satisfying answers to many relevant questions in radiation therapy. As they still fall short of consistently providing correct information, it is problematic to use them for obtaining medical information. As LLMs will further improve in the future, they are expected to have an increasing impact not only on general society, but also on clinical practice, including radiation oncology.
- Achieving consensus, coherence, clarity and consistency when talking about addictionRobert West, Sharon Cox, Caitlin Jade Notley, Guy Du Plessis, and Janna HastingsAddiction, Nov 2023
- AI for Scientific DiscoveryJanna HastingsJun 2023
AI for Scientific Discovery provides an accessible introduction to the wide-ranging applications of artificial intelligence (AI) technologies in scientific research and discovery across the full breadth of scientific disciplines. AI technologies support discovery science in multiple ways. They support literature management and synthesis, allowing the wealth of what has already been discovered and reported on to be integrated and easily accessed. They play a central role in data analysis and interpretation in the context of what is called ‘data science’. AI is also helping to combat the reproducibility crisis in scientific research by underpinning the discovery process with AI-enabled standards and pipelines and supporting the management of large-scale data and knowledge resources so that they can be shared and integrated and serve as a background ‘knowledge ecosystem’ into which new discoveries can be embedded. However, there are limitations to what AI can achieve and its outputs can be biased and confounded and thus should not be blindly trusted. The latest generation of hybrid and ‘human-in-the-loop’ AI technologies have as their objective a balance between human inputs and insights and the power of number-crunching and statistical inference at a massive scale that AI technologies are best at.
- Applied ontology: Where are we now and where are we going?Janna Hastings and John A. BatemanApplied Ontology, Jan 2023
- The Behaviour Change Technique Ontology: Transforming the Behaviour Change Technique Taxonomy v1Marta M. Marques, Alison J. Wright, Elizabeth Corker, Marie Johnston, Robert West, Janna Hastings, Lisa Zhang, and Susan MichieWellcome Open Research, Jul 2023
Background: The Behaviour Change Technique Taxonomy v1 (BCTTv1) specifies the potentially active content of behaviour change interventions. Evaluation of BCTTv1 showed the need to extend it into a formal ontology, improve its labels and definitions, add BCTs and subdivide existing BCTs. We aimed to develop a Behaviour Change Technique Ontology (BCTO) that would meet these needs., Methods: The BCTO was developed by: (1) collating and synthesising feedback from multiple sources; (2) extracting information from published studies and classification systems; (3) multiple iterations of reviewing and refining entities, and their labels, definitions and relationships; (4) refining the ontology via expert stakeholder review of its comprehensiveness and clarity; (5) testing whether researchers could reliably apply the ontology to identify BCTs in intervention reports; and (6) making it available online and creating a machine-readable version., Results: Initially there were 282 proposed changes to BCTTv1. Following first-round review, 19 BCTs were split into two or more BCTs, 27 new BCTs were added and 26 BCTs were moved into a different group, giving 161 BCTs hierarchically organised into 12 logically defined higher-level groups in up to five hierarchical levels. Following expert stakeholder review, the refined ontology had 247 BCTs hierarchically organised into 20 higher-level groups. Independent annotations of intervention evaluation reports by researchers familiar and unfamiliar with the ontology resulted in good levels of inter-rater reliability (0.82 and 0.79, respectively). Following revision informed by this exercise, 34 BCTs were added, resulting in a final version of the BCTO containing 281 BCTs organised into 20 higher-level groups over five hierarchical levels., Discussion: The BCT Ontology provides a standard terminology and comprehensive classification system for the content of behaviour change interventions that can be reliably used to describe interventions.
- Ontologies of behaviour: Current perspectives and future potential in health psychologyHarriet M. Baird, Janna Hastings, Marie Johnston, Susan F. Michie, Paul Norman, Gabriel Nudelman, Alexander J. Scott, Thomas Llewelyn Webb, Robert West, and Alison J. WrightEuropean Health Psychologist, Jul 2023
- Knowledge Graphs for the Life Sciences: Recent Developments, Challenges and OpportunitiesJiaoyan Chen, Hang Dong, Janna Hastings, Ernesto Jiménez-Ruiz, Vanessa López, Pierre Monnin, Catia Pesquita, Petr Škoda, and Valentina TammaTransactions on Graph Data and Knowledge, Jul 2023
The term life sciences refers to the disciplines that study living organisms and life processes, and include chemistry, biology, medicine, and a range of other related disciplines. Research efforts in life sciences are heavily data-driven, as they produce and consume vast amounts of scientific data, much of which is intrinsically relational and graph-structured. The volume of data and the complexity of scientific concepts and relations referred to therein promote the application of advanced knowledge-driven technologies for managing and interpreting data, with the ultimate aim to advance scientific discovery. In this survey and position paper, we discuss recent developments and advances in the use of graph-based technologies in life sciences and set out a vision for how these technologies will impact these fields into the future. We focus on three broad topics: the construction and management of Knowledge Graphs (KGs), the use of KGs and associated technologies in the discovery of new knowledge, and the use of KGs in artificial intelligence applications to support explanations (explainable AI). We select a few exemplary use cases for each topic, discuss the challenges and open research questions within these topics, and conclude with a perspective and outlook that summarizes the overarching challenges and their potential solutions as a guide for future research.
2022
- Interpretable Ontology Extension in ChemistryMartin Glauer, Adel Memariani, Fabian Neuhaus, Till Mossakowski, and Janna HastingsSemantic Web Journal, Feb 2022
We present an approach towards ontology extension that uses structural information to train a transformer-based model that predicts new subsumption relations. The ELECTRA model has been pre-trained using a combination of molecules from the ChEBI ontology and a selection of molecules from the PubChem database (chebai/data/SWJpre/raw/smiles.txt). The resulting model has then been fine-truned on a selection of ChEBI classes. The trained model has then been applied to a set of previously unseen chemicals from PubChem (hazardous.txt). The resulting predictions have been used to extend the ChEBI ontology. The extended ontology can be found as an owl file in ’chebi-slim-extended.owl.gz’ and as a plot in ’classif-hazardous.png’. The resulting ontology was inconsistent because some of the predicted subsumption relations violated disjointness axioms. Those subsumption relations have been removed (’chebi-slim-extended-fixed.owl.gz’). The README.md file describes how to reproduce our results.
- More phenomenology in psychiatry? Applied ontology as a method towards integrationRasmus R Larsen, Luca F Maschião, Valter L Piedade, Guilherme Messas, and Janna HastingsThe Lancet Psychiatry, Sep 2022
There have been renewed calls to use phenomenology in psychiatry to improve knowledge about causation, diagnostics, and treatment of mental health conditions. A phenomenological approach aims to elucidate the subjective experiences of mental health, which its advocates claim have been largely neglected by current diagnostic frameworks in psychiatry (eg, DSM-5). The consequence of neglecting rich phenomenological information is a comparatively more constrained approach to theory development, empirical research, and care programmes. Although calls for more phenomenology in psychiatry have been met with enthusiasm, there is still relatively little information on how to practically facilitate this integration. In this Personal View, we argue that phenomenological approaches need a shared semantic framework to drive their innovative potential, thus enabling consistent data capture, exchange, and interoperability with current mental health data and informatics approaches (eg, the Research Domain Criteria project). We show how an applied ontology of phenomenological psychopathology offers a suitable method to address these challenges.
- ESC-Rules: Explainable, Semantically Constrained Rule SetsMartin Glauer, Robert West, Susan Michie, and Janna HastingsAug 2022
We describe a novel approach to explainable prediction of a continuous variable based on learning fuzzy weighted rules. Our model trains a set of weighted rules to maximise prediction accuracy and minimise an ontology-based ’semantic loss’ function including user-specified constraints on the rules that should be learned in order to maximise the explainability of the resulting rule set from a user perspective. This system fuses quantitative sub-symbolic learning with symbolic learning and constraints based on domain knowledge. We illustrate our system on a case study in predicting the outcomes of behavioural interventions for smoking cessation, and show that it outperforms other interpretable approaches, achieving performance close to that of a deep learning model, while offering transparent explainability that is an essential requirement for decision-makers in the health domain.
- Ontology development is consensus creation, not (merely) representationFabian Neuhaus and Janna HastingsApplied Ontology, Jan 2022
Ontology development methodologies emphasise knowledge gathering from domain experts and documentary resources, and knowledge representation using an ontology language such as OWL or FOL. However, working ontologists are often surprised by how challe
- Achieving Inclusivity by Design: Social and Contextual Information in Medical KnowledgeJanna HastingsYearbook of Medical Informatics, Jun 2022
Objectives: To select, present, and summarize the most relevant papers published in 2020 and 2021 in the field of Knowledge Representation and Knowledge Management, Medical Vocabularies and Ontologies, with a particular focus on health inclusivity and bias. Methods: A broad search of the medical literature indexed in PubMed was conducted. The search terms ’ontology’/’ontologies’ or ’medical knowledge management’ for the dates 2020-2021 (search conducted November 26, 2021) returned 9,608 records. These were pre-screened based on a review of the titles for relevance to health inclusivity, bias, social and contextual factors, and health behaviours. Among these, 109 papers were selected for in-depth reviewing based on full text, from which 22 were selected for inclusion in this survey. Results: Selected papers were grouped into three themes, each addressing one aspect of the overall challenge for medical knowledge management. The first theme addressed the development of ontologies for social and contextual factors broadening the scope of health information. The second theme addressed the need for synthesis and translation of knowledge across historical disciplinary boundaries to address inequities and bias. The third theme encompassed a growing interest in the semantics of datasets used to train medical artificial intelligence systems and on how to ensure they are free of bias. Conclusions: Medical knowledge management and semantic resources have much to offer efforts to tackle bias and enhance health inclusivity. Tackling inequities and biases requires relevant, semantically rich data, which needs to be captured and exchanged.
2021
- Why and how to engage expert stakeholders in ontology development: insights from social and behavioural sciencesEmma Norris, Janna Hastings, Marta M. Marques, Ailbhe N. Finnerty Mutlu, Silje Zink, and Susan MichieJournal of Biomedical Semantics, Mar 2021
Incorporating the feedback of expert stakeholders in ontology development is important to ensure content is appropriate, comprehensive, meets community needs and is interoperable with other ontologies and classification systems. However, domain experts are often not formally engaged in ontology development, and there is little available guidance on how this involvement should best be conducted and managed. Social and behavioural science studies often involve expert feedback in the development of tools and classification systems but have had little engagement with ontology development. This paper aims to (i) demonstrate how expert feedback can enhance ontology development, and (ii) provide practical recommendations on how to conduct expert feedback in ontology development using methodologies from the social and behavioural sciences.
- Informatics Technologies for the Acquisition of Psychological, Behavioral, Interpersonal, Social and Environmental DataElena Tenenbaum, Piper A. Ranallo, and Janna HastingsIn Mental Health Informatics: Enabling a Learning Mental Healthcare System, Mar 2021
The collection, capture, storage, sharing, and interpretation of data are essential to all research and practice in mental health. A wide range of informatics technologies and tools have been developed to facilitate data use across the full knowledge acquisition lifecycle. In this chapter, we focus on data acquisition. We introduce the field of psychometrics—the science of measurement in psychology. We discuss the unique challenges of data acquisition in mental health by exploring the nature of psychological, behavioral, interpersonal, social, and environmental data in the context of mental health. Finally, we discuss some current challenges in the use of informatics technologies for data in these domains and how those challenges might be addressed in the future.
- Nuclear and cytoplasmic huntingtin inclusions exhibit distinct biochemical composition, interactome and ultrastructural propertiesNathan Riguet, Anne-Laure Mahul-Mellier, Niran Maharjan, Johannes Burtscher, Marie Croisier, Graham Knott, Janna Hastings, Alice Patin, Veronika Reiterer, Hesso Farhan, and 2 more authorsNature Communications, Nov 2021
Despite the strong evidence linking the aggregation of the Huntingtin protein (Htt) to the pathogenesis of Huntington’s disease (HD), the mechanisms underlying Htt aggregation and neurodegeneration remain poorly understood. Herein, we investigated the ultrastructural properties and protein composition of Htt cytoplasmic and nuclear inclusions in mammalian cells and primary neurons overexpressing mutant exon1 of the Htt protein. Our findings provide unique insight into the ultrastructural properties of cytoplasmic and nuclear Htt inclusions and their mechanisms of formation. We show that Htt inclusion formation and maturation are complex processes that, although initially driven by polyQ-dependent Htt aggregation, also involve the polyQ and PRD domain-dependent sequestration of lipids and cytoplasmic and cytoskeletal proteins related to HD dysregulated pathways; the recruitment and accumulation of remodeled or dysfunctional membranous organelles, and the impairment of the protein quality control and degradation machinery. We also show that nuclear and cytoplasmic Htt inclusions exhibit distinct biochemical compositions and ultrastructural properties, suggesting different mechanisms of aggregation and toxicity.
- Scientific Ontologies, Digital Curation and the Learning Knowledge EcosystemJanna HastingsQurator, Nov 2021
The global coronavirus pandemic has brought another ongoing crisis into the spotlight: that of digital misinformation. While society at a global scale is facing challenges that demand scientific solutions as never before, trust in experts and scientific expertise is falling, and conspiracy theories abound.
- Introducing the Open Energy Ontology: Enhancing data interpretation and interfacing in energy systems analysisMeisam Booshehri, Lukas Emele, Simon Flügel, Hannah Förster, Johannes Frey, Ulrich Frey, Martin Glauer, Janna Hastings, Christian Hofmann, Carsten Hoyer-Klick, and 12 more authorsEnergy and AI, Sep 2021
Heterogeneous data, different definitions and incompatible models are a huge problem in many domains, with no exception for the field of energy systems analysis. Hence, it is hard to re-use results, compare model results or couple models at all. Ontologies provide a precisely defined vocabulary to build a common and shared conceptualisation of the energy domain. Here, we present the Open Energy Ontology (OEO) developed for the domain of energy systems analysis. Using the OEO provides several benefits for the community. First, it enables consistent annotation of large amounts of data from various research projects. One example is the Open Energy Platform (OEP). Adding such annotations makes data semantically searchable, exchangeable, re-usable and interoperable. Second, computational model coupling becomes much easier. The advantages of using an ontology such as the OEO are demonstrated with three use cases: data representation, data annotation and interface homogenisation. We also describe how the ontology can be used for linked open data (LOD).
- Non-monotonic fibril surface occlusion by GFP tags from coarse-grained molecular simulationsJulian C. Shillcock, Janna Hastings, Nathan Riguet, and Hilal A. LashuelComputational and Structural Biotechnology Journal, Dec 2021
The pathological growth of amyloid fibrils in neurons underlies the progression of neurodegenerative diseases including Alzheimer’s and Parkinson’s disease. Fibrils form when soluble monomers oligomerise in the cytoplasm. Their subsequent growth occurs via nucleated polymerization mechanisms involving the free ends of the fibrils augmented by secondary nucleation of new oligomers at their surface. Amyloid fibrils possess a complex interactome with diffusing cytoplasmic proteins that regulates many aspects of their growth, seeding capacity, biochemical activity and transition to pathological inclusions in diseased brains. Changes to their surface are also expected to modify their interactome, pathogenicity and spreading in the brain. Many assays visualise fibril formation, growth and inclusion formation by decorating monomeric proteins with fluorescent tags such as GFP. Recent studies from our group suggest that tags with sizes comparable to the fibril radius may modify the fibril surface accessibility and thus their PTM pattern, interactome and ability to form inclusions. Using coarse-grained molecular simulations of a single alpha synuclein fibril tagged with GFP we find that thermal fluctuations of the tags create a non-monotonic, size-dependent sieve around the fibril that perturbs its interactome with diffusing species. Our results indicate that experiments using tagged and untagged monomers to study the growth and interactome of fibrils should be compared with caution, and the confounding effects of the tags are more complex than a reduction in surface accessibility. The prevalence of fluorescent tags in amyloid fibril growth experiments suggests this has implications beyond the specific alpha synuclein fibrils we model here.
- Automated and Explainable Ontology Extension Based on Deep Learning: A Case Study in the Chemical DomainAdel Memariani, Martin Glauer, Fabian Neuhaus, Till Mossakowski, and Janna HastingsarXiv:2109.09202 [cs], Sep 2021
Reference ontologies provide a shared vocabulary and knowledge resource for their domain. Manual construction enables them to maintain a high quality, allowing them to be widely accepted across their community. However, the manual development process does not scale for large domains. We present a new methodology for automatic ontology extension and apply it to the ChEBI ontology, a prominent reference ontology for life sciences chemistry. We trained a Transformer-based deep learning model on the leaf node structures from the ChEBI ontology and the classes to which they belong. The model is then capable of automatically classifying previously unseen chemical structures. The proposed model achieved an overall F1 score of 0.80, an improvement of 6 percentage points over our previous results on the same dataset. Additionally, we demonstrate how visualizing the model’s attention weights can help to explain the results by providing insight into how the model made its decisions.
- FOWL – An OWL to FOL TranslatorSimon Flügel, Anna Kleinau, Fabian Neuhaus, Martin Glauer, and Janna HastingsIn Proceedings of the Formal Ontology in Information Systems conference, Bolzano, Italy, Sep 2021
The Web Ontology Language OWL 2 DL is the language used to build ontologies for a wide range of domains including the life sciences, the financial industry and the domain of energy simulations. Many of these domain ontologies are built beneath upper ontologies (such as DOLCE or BFO), which themselves are written in more expressive logics than OWL 2 DL, in particular first-order logic (although they may provide truncated versions in OWL). In this paper we present FOWL, a tool that enables the translation of OWL ontologies into FOL, and thereby the integration of OWL domain ontologies with first-order logic ontologies. One use case for this tool is the ability to validate OWL domain ontologies against their upper-level ontology. FOWL also allows reasoning over the translation using the VAMPIRE reasoner.
- Specifying who delivers behaviour change interventions: development of an Intervention Source OntologyEmma Norris, Alison J. Wright, Janna Hastings, Robert West, Neil Boyt, and Susan MichieApr 2021
Background: Identifying how behaviour change interventions are delivered, including by whom, is key to understanding intervention effectiveness. However, information about who delivers interventions is reported inconsistently in intervention evaluations, limiting communication and knowledge accumulation. This paper reports a method for consistent reporting: The Intervention Source Ontology. This forms one part of the Behaviour Change Intervention Ontology, which aims to cover all aspects of behaviour change interventions . Methods : The Intervention Source Ontology was developed following methods for ontology development and maintenance used in the Human Behaviour-Change Project, with seven key steps: 1) define the scope of the ontology, 2) identify key entities and develop their preliminary definitions by reviewing existing classification systems (top-down) and reviewing 100 behaviour change intervention reports (bottom-up), 3) refine the ontology by piloting the preliminary ontology on 100 reports, 4) stakeholder review by 34 behavioural science and public health experts, 5) inter-rater reliability testing of annotating intervention reports using the ontology, 6) specify ontological relationships between entities and 7) disseminate and maintain the Intervention Source Ontology. Results: The Intervention Source Ontology consists of 140 entities. Key areas of the ontology include Occupational Role of Source , Relatedness between Person Source and the Target Population , Sociodemographic attributes and Expertise. Inter-rater reliability was found to be 0.60 for those familiar with the ontology and 0.59 for those unfamiliar with it, levels of agreement considered ‘acceptable’. Conclusions: Information about who delivers behaviour change interventions can be reliably specified using the Intervention Source Ontology. For human-delivered interventions, the ontology can be used to classify source characteristics in existing behaviour change reports and enable clearer specification of intervention sources in reporting.
- Integrative Paradigms for Knowledge Discovery in Mental Health: Overcoming the Fragmentation of Knowledge Inherent in Disparate Theoretical ParadigmsJanna Hastings and Rasmus Rosenberg LarsenIn Mental Health Informatics: Enabling a Learning Mental Healthcare System, Sep 2021
The domain of mental health is inherently complex, spanning across multiple disciplines, data types, descriptive levels, and approaches. This complexity has brought considerable challenges in terms of how to facilitate efficient knowledge discovery and integration across disciplines in the domain. The vocabulary and semantic frameworks in use across these different descriptive levels are fragmented and contested, and it is difficult to gain an overview of what is known across all the relevant bodies of knowledge and practice. In this chapter, we review progress that has recently been made towards integrative semantic and computational frameworks for structuring and advancing mental health research. This includes the paradigm shift incubated in the NIMH’s RDoC effort, which offers a roadmap for studying the nature of the complex interactions within and between human systems: biological (body, brain), mental (mind), behavioral, social, and environmental. We also review computational approaches to infer and model relationships between entities that explicitly cross levels of explanation and disciplinary boundaries. We describe the quantitative methods that are used to integrate and analyze across heterogeneous datasets, and the epistemological challenges that face the field when attempting to determine mechanistic explanations that move the global understanding of mental health forward.
- Learning chemistry: exploring the suitability of machine learning for the task of structure-based chemical ontology classificationJanna Hastings, Martin Glauer, Adel Memariani, Fabian Neuhaus, and Till MossakowskiJournal of Cheminformatics, Mar 2021
Chemical data is increasingly openly available in databases such as PubChem, which contains approximately 110 million compound entries as of February 2021. With the availability of data at such scale, the burden has shifted to organisation, analysis and interpretation. Chemical ontologies provide structured classifications of chemical entities that can be used for navigation and filtering of the large chemical space. ChEBI is a prominent example of a chemical ontology, widely used in life science contexts. However, ChEBI is manually maintained and as such cannot easily scale to the full scope of public chemical data. There is a need for tools that are able to automatically classify chemical data into chemical ontologies, which can be framed as a hierarchical multi-class classification problem. In this paper we evaluate machine learning approaches for this task, comparing different learning frameworks including logistic regression, decision trees and long short-term memory artificial neural networks, and different encoding approaches for the chemical structures, including cheminformatics fingerprints and character-based encoding from chemical line notation representations. We find that classical learning approaches such as logistic regression perform well with sets of relatively specific, disjoint chemical classes, while the neural network is able to handle larger sets of overlapping classes but needs more examples per class to learn from, and is not able to make a class prediction for every molecule. Future work will explore hybrid and ensemble approaches, as well as alternative network architectures including neuro-symbolic approaches.
2020
- Theory and ontology in behavioural scienceJanna Hastings, Susan Michie, and Marie JohnstonNature Human Behaviour, Mar 2020
- An ontology-based modelling system (OBMS) for representing behaviour change theories applied to 76 theoriesJoanna Hale, Janna Hastings, Robert West, Carmen E. Lefevre, Artur Direito, Lauren Connell Bohlen, Cristina Godinho, Niall Anderson, Silje Zink, Hilary Groarke, and 1 more authorWellcome Open Research, Jul 2020
Background: To efficiently search, compare, test and integrate behaviour change theories, they need to be specified in a way that is clear, consistent and computable. An ontology-based modelling system (OBMS) has previously been shown to be able to represent five commonly used theories in this way. We aimed to assess whether the OBMS could be applied more widely and to create a database of behaviour change theories, their constructs and propositions. Methods: We labelled the constructs within 71 theories and used the OBMS to represent the relationships between the constructs. Diagrams of each theory were sent to authors or experts for feedback and amendment. The 71 finalised diagrams plus the five previously generated diagrams were used to create a searchable database of 76 theories in the form of construct-relationship-construct triples. We conducted a set of illustrative analyses to characterise theories in the database. Results: All 71 theories could be satisfactorily represented using this system. In total, 35 (49%) were finalised with no or very minor amendment. The remaining 36 (51%) were finalised after changes to the constructs (seven theories), relationships between constructs (15 theories) or both (14 theories) following author/expert feedback. The mean number of constructs per theory was 20 (min. = 6, max. = 72), with the mean number of triples per theory 31 (min. = 7, max. = 89). Fourteen distinct relationship types were used, of which the most commonly used was ‘influences’, followed by ‘part of’. Conclusions: The OBMS can represent a wide array of behavioural theories in a precise, computable format. This system should provide a basis for better integration and synthesis of theories than has hitherto been possible.
- Representation of behaviour change interventions and their evaluation: Development of the Upper Level of the Behaviour Change Intervention OntologySusan Michie, Robert West, Ailbhe N. Finnerty, Emma Norris, Alison J. Wright, Marta M. Marques, Marie Johnston, Michael P. Kelly, James Thomas, and Janna HastingsWellcome Open Research, Jun 2020
Background: Behaviour change interventions (BCI), their contexts and evaluation methods are heterogeneous, making it difficult to synthesise evidence and make recommendations for real-world policy and practice. Ontologies provide a means for addressing this. They represent knowledge formally as entities and relationships using a common language able to cross disciplinary boundaries and topic domains. This paper reports the development of the upper level of the Behaviour Change Intervention Ontology (BCIO), which provides a systematic way to characterise BCIs, their contexts and their evaluations. Methods: Development took place in four steps. (1) Entities and relationships were identified by behavioural and social science experts, based on their knowledge of evidence and theory, and their practical experience of behaviour change interventions and evaluations. (2) The outputs of the first step were critically examined by a wider group of experts, including the study ontology expert and those experienced in annotating relevant literature using the initial ontology entities. The outputs of the second step were tested by (3) feedback from three external international experts in ontologies and (4) application of the prototype upper-level BCIO to annotating published reports; this informed the final development of the upper-level BCIO. Results: The final upper-level BCIO specifies 42 entities, including the BCI scenario, elaborated across 21 entities and 7 relationship types, and the BCI evaluation study comprising 10 entities and 9 relationship types. BCI scenario entities include the behaviour change intervention (content and delivery), outcome behaviour, mechanism of action, and its context, which includes population and setting. These entities have corresponding entities relating to the planning and reporting of interventions and their evaluations. Conclusions: The upper level of the BCIO provides a comprehensive and systematic framework for representing BCIs, their contexts and their evaluations.
- Delivering Behaviour Change Interventions: Development of a Mode of Delivery OntologyMarta M. Marques, Rachel N. Carey, Emma Norris, Fiona Evans, Ailbhe N. Finnerty, Janna Hastings, Ella Jenkins, Marie Johnston, Robert West, and Susan MichieWellcome Open Research, Jun 2020
Background: Investigating and improving the effects of behaviour change interventions requires detailed and consistent specification of all aspects of interventions. An important feature of interventions is the way in which these are delivered, i.e. their mode of delivery. This paper describes an ontology for specifying the mode of delivery of interventions, which forms part of the Behaviour Change Intervention Ontology, currently being developed in the Wellcome Trust funded Human Behaviour-Change Project. Methods: The Mode of Delivery Ontology was developed in an iterative process of annotating behaviour change interventions evaluation reports, and consulting with expert stakeholders. It consisted of seven steps: 1) annotation of 110 intervention reports to develop a preliminary classification of modes of delivery; 2) open review from international experts (n=25); 3) second round of annotations with 55 reports to test inter-rater reliability and identify limitations; 4) second round of expert review feedback (n=16); 5) final round of testing of the refined ontology by two annotators familiar and two annotators unfamiliar with the ontology; 6) specification of ontological relationships between entities; and 7) transformation into a machine-readable format using the Web Ontology Language (OWL) language and publishing online. Results: The resulting ontology is a four-level hierarchical structure comprising 65 unique modes of delivery, organised by 15 upper-level classes: Informational , Environmental change, Somatic, Somatic alteration, Individual-based/ Pair-based /Group-based, Uni-directional/Interactional, Synchronous/ Asynchronous, Push/ Pull, Gamification, Arts feature. Relationships between entities consist of is_a . Inter-rater reliability of the Mode of Delivery Ontology for annotating intervention evaluation reports was a =0.80 (very good) for those familiar with the ontology and a = 0.58 (acceptable) for those unfamiliar with it. Conclusion: The ontology can be used for both annotating and writing behaviour change intervention evaluation reports in a consistent and coherent manner, thereby improving evidence comparison, synthesis, replication, and implementation of effective interventions.
- The Human Behaviour-Change Project: An artificial intelligence system to answer questions about changing behaviourSusan Michie, James Thomas, Pol Mac Aonghusa, Robert West, Marie Johnston, Michael P. Kelly, John Shawe-Taylor, Janna Hastings, Francesca Bonin, and Alison O’Mara-EvesWellcome Open Research, Jun 2020
Changing behaviour is necessary to address many of the threats facing human populations. However, identifying behaviour change interventions likely to be effective in particular contexts as a basis for improving them presents a major challenge. The Human Behaviour-Change Project harnesses the power of artificial intelligence and behavioural science to organise global evidence about behaviour change to predict outcomes in common and unknown behaviour change scenarios.
- Ontologies relevant to behaviour change interventions: a method for their developmentAlison J. Wright, Emma Norris, Ailbhe N. Finnerty, Marta M. Marques, Marie Johnston, Michael P. Kelly, Janna Hastings, Robert West, and Susan MichieWellcome Open Research, Aug 2020
Background: Behaviour and behaviour change are integral to many aspects of wellbeing and sustainability. However, reporting behaviour change interventions accurately and synthesising evidence about effective interventions is hindered by lacking a shared, scientific terminology to describe intervention characteristics. Ontologies are knowledge structures that provide controlled vocabularies to help unify and connect scientific fields. To date, there is no published guidance on the specific methods required to develop ontologies relevant to behaviour change. We report the creation and refinement of a method for developing ontologies that make up the Behaviour Change Intervention Ontology (BCIO). Aims: (1) To describe the development method of the BCIO and explain its rationale; (2) To provide guidance on implementing the activities within the development method. Method and results: The method for developing ontologies relevant to behaviour change interventions was constructed by considering principles of good practice in ontology development and identifying key activities required to follow those principles. The method’s details were refined through application to developing two ontologies. The resulting ontology development method involved: (1) defining the ontology’s scope; (2) identifying key entities; (3) refining the ontology through an iterative process of literature annotation, discussion and revision; (4) expert stakeholder review; (5) testing inter-rater reliability; (6) specifying relationships between entities, and; (7) disseminating and maintaining the ontology. Guidance is provided for conducting relevant activities for each step. Conclusions: We have developed a detailed method for creating ontologies relevant to behaviour change interventions, together with practical guidance for each step, reflecting principles of good practice in ontology development. The most novel aspects of the method are the use of formal mechanisms for literature annotation and expert stakeholder review to develop and improve the ontology content. We suggest the mnemonic SELAR3, representing the method’s first six steps as Scope, Entities, Literature Annotation, Review, Reliability, Relationships.
- The case for development of an E-cigarette Ontology (E-CigO) to improve quality, efficiency and clarity in the conduct and interpretation of researchSharon Cox, Janna Hastings, Robert West, and Caitlin NotleyQeios, Apr 2020
There is an urgent need for more clarity and consistency in the use of terms in the field of e-cigarette research. Ontologies are computer artifacts that are increasingly widely used in science to represent knowledge in terms of uniquely defined entities and their relationships with other entities. These are constructed in a way that promotes clarity of thinking, much more effective searching, inference and interoperability across domains of study. We are constructing an E-cigarette Ontology (ECig-O) covering all the types of entity that are referred to in reports of e-cigarette research. It is part a larger Addiction Ontology (AddictO) covering all aspects of addiction.
- Addiction Ontology: Applying Basic Formal Ontology in the Addiction domainJanna Hastings, Sharon Cox, Robert West, and Caitlin NotleyQeios, Dec 2020
\textlessdiv class="ck-publication-abstract-context"\textgreater\textlessp\textgreaterOntologies are being used in many areas of science to improve clarity and communication of research methods, findings and theories. Many of these ontologies use an upper level ontology called Basic Formal Ontology (BFO) as their frame of reference. This article summarises Basic Formal Ontology and shows how it can provide a basis for development of an Addiction Ontology that encompasses all the things that addiction researchers, practitioners and policy makers want to refer to. BFO makes a fundamental distinction between what it calls continuants (e.g. objects and their characteristics) and occurrents (e.g. processes). Classifying addiction-related entities using this system enables important distinctions to be made that are frequently overlooked or confused in the literature due to inherent ambiguities in natural language expressions. The Addiction Ontology uses this framework to convey information about: people and populations and their characteristics (e.g. substance use disorder), products (e.g. heroin, tobacco-containing products), behaviours (e.g. cigarette smoking, alcohol consumption), interventions (e.g. detoxification, rehabilitation, legislation), research (e.g. measurement, theories, study designs), organisations (e.g. pharmaceutical industry, tobacco companies), and settings (e.g. hospital outpatient clinic, country).\textless/p\textgreater\textless/div\textgreater
- Using Genome-Scale Metabolic Networks for Analysis, Visualization, and Integration of Targeted Metabolomics DataJake P. N. Hattwell, Janna Hastings, Olivia Casanueva, Horst Joachim Schirra, and Michael WittingIn Computational Methods and Data Analysis for Metabolomics, Dec 2020
Interpretation of metabolomics data in the context of biological pathways is important to gain knowledge about underlying metabolic processes. In this chapter we present methods to analyze genome-scale models (GSMs) and metabolomics data together. This includes reading and mining of GSMs using the SBTab format to retrieve information on genes, reactions, and metabolites. Furthermore, the chapter showcases the generation of metabolic pathway maps using the Escher tool, which can be used for data visualization. Lastly, approaches to constrain flux balance analysis (FBA) by metabolomics data are presented.
2019
- Flow with the flux: Systems biology tools predict metabolic drivers of ageing in C. elegansJanna Hastings, Manusnan Suriyalaksh, and Olivia CasanuevaCurrent Opinion in Systems Biology, Feb 2019
Ageing was thought to be an immutable process, but over the past few decades, many environmental and genetic perturbations have been shown to dramatically alter lifespan. This plasticity, however, is not normally used, and an early decline in key cellular functions such as stress responses and metabolism can cause late-life pathologies. To extend healthspan, it is therefore essential to identify the earliest molecular changes and determine their contribution to late-life pathologies. Caenorhabditis elegans is an ideal model organism to perform detailed temporally resolved multi-omics measurements. Transomics data can be harnessed for mechanistic systems biology tools such as flux balance and network inference analyses that are able to disentangle cause from effect during ageing.
- Addiction Theories and Constructs: a new series.Robert West, John Marsden, and Janna HastingsAddiction (Abingdon, England), Jun 2019
- The Literary Theme Ontology for Media Annotation and Information RetrievalPaul Sheridan, Mikael Onsjö, and Janna HastingsarXiv:1905.00522 [cs], Aug 2019
Literary theme identification and interpretation is a focal point of literary studies scholarship. Classical forms of literary scholarship, such as close reading, have flourished with scarcely any need for commonly defined literary themes. However, the rise in popularity of collaborative and algorithmic analyses of literary themes in works of fiction, together with a requirement for computational searching and indexing facilities for large corpora, creates the need for a collection of shared literary themes to ensure common terminology and definitions. To address this need, we here introduce a first draft of the Literary Theme Ontology. Inspired by a traditional framing from literary theory, the ontology comprises literary themes drawn from the authors own analyses, reference books, and online sources. The ontology is available at https://github.com/theme-ontology/lto under a Creative Commons Attribution 4.0 International license (CC BY 4.0).
- Development of a formal system for representing behaviour-change theoriesRobert West, Cristina A. Godinho, Lauren Connell Bohlen, Rachel N. Carey, Janna Hastings, Carmen E. Lefevre, and Susan MichieNature Human Behaviour, May 2019
Use of natural language to represent behaviour-change theories has resulted in lack of clarity and consistency, hindering comparison, integration, development and use. This paper describes development of a formal system for representing behaviour-change theories that aims to improve clarity and consistency. A given theory is represented in terms of (1) its component constructs (for example, ‘self-efficacy’, ‘perceived threat’ or ‘subjective norm’), which are labelled and defined, and (2) relationships between pairs of constructs, which may be causal, structural or semantic. This formalism appears adequate to represent five commonly used theories (health belief model, information–motivation–behavioural skill model, social cognitive theory, theory of planned behaviour and the trans-theoretical model). Theory authors and experts judged that the system was able to capture the main propositions of the theories. Following this proof of concept, the next step is to assess how far the system can be applied to other theories of behaviour change.
- A scoping review of ontologies related to human behaviour changeEmma Norris, Ailbhe N. Finnerty, Janna Hastings, Gillian Stokes, and Susan MichieNature Human Behaviour, Feb 2019
Ontologies are classification systems specifying entities, definitions and inter-relationships for a given domain, with the potential to advance knowledge about human behaviour change. A scoping review was conducted to: (1) identify what ontologies exist related to human behaviour change, (2) describe the methods used to develop these ontologies and (3) assess the quality of identified ontologies. Using a systematic search, 2,303 papers were identified. Fifteen ontologies met the eligibility criteria for inclusion, developed in areas such as cognition, mental disease and emotions. Methods used for developing the ontologies were expert consultation, data-driven techniques and reuse of terms from existing taxonomies, terminologies and ontologies. Best practices used in ontology development and maintenance were documented. The review did not identify any ontologies representing the breadth and detail of human behaviour change. This suggests that advancing behavioural science would benefit from the development of a behaviour change intervention ontology.
- Representing Literary Characters and their Attributes in an OntologyJanna Hastings and Stefan SchulzProceedings of the Joint Ontology Workshops (JOWO), Feb 2019
The digital humanities is a burgeoning field of research, bringing computational methods to literary investigations. Ontologies are computational structures that contain descriptions of entities and relations in a domain, and as such they form natural hubs for indexing, search and retrieval as well as enable sophisticated automated inferencing applications. Ontologies are widely used in the medical and biological domains, and are beginning to be adopted in other disciplines such as the social sciences. However, there are unique challenges with the adoption of ontologies for the representation of the elements of literary works. Literary meanings are subjective to some extent, and the ontological mode of being of fictional entities is different from that of the entities in the real world that are the subject matter of the sciences. We focus here on elucidating these challenges through the lens of fictional characters (such as, for example, Macbeth) and the social and personal attributes they are described as having (e.g. age and nationality). We develop a detailed representational strategy – a pattern – for representing fictional characters and their attributes in OWL, in applied ontologies.
- Theory and Ontology in Building Cumulative Behavioural ScienceJanna Hastings, Susan Michie, and Marie JohnstonFeb 2019
The robust use of theory as a driver for research and evidence synthesis has the potential to mitigate the reproducibility crisis and contribute to the accumulation of knowledge and progress in the field of behavioural science. However, agreement on a single theory or theoretical framework is highly unlikely and arguably undesirable. We suggest that an alternative approach is grounded in the use of ontologies: formal computational structures for clearly defining entities and relations that enable theoretical integration via a network of relations between entities described in different theories. Ontologies are already widely adopted in the life sciences, but as yet have seen little adoption in the behavioural sciences. T hey have the potential both for comparison between theories, and for aggregation and evidence synthesis regardless of the theoretical framework that led to the generation of findings, leading to genuine cumulative progress
2018
- Modeling meets Metabolomics – The WormJam Consensus Model as basis for Metabolic Studies in the model organism \textitCaenorhabditis elegansMichael Witting, Janna Hastings, Nicolas Rodriguez, Chintan J. Joshi, Jake P. Hattwell, Paul R. Ebert, Michel van Weeghel, Michael Wakelam, Riekelt Houtkooper, Abraham Mains, and 7 more authorsFrontiers in Molecular Biosciences, Feb 2018
- From Affective Science to Psychiatric Disorder: Ontology as a Semantic BridgeRasmus Rosenberg Larsen and Janna HastingsFrontiers in Psychiatry, Feb 2018
Advances in emotion and affective science have yet to translate routinely into psychiatric research and practice. This is unfortunate since emotion and affect are fundamental components of many psychiatric conditions. Rectifying this lack of interdisciplinary integration could thus be a potential avenue for improving psychiatric diagnosis and treatment. In this contribution, we propose and discuss an ontological framework for explicitly capturing the complex interrelations between affective entities and psychiatric disorders, in order to facilitate mapping and integration between affective science and psychiatric diagnostics. We build on and enhance the categorisation of emotion, affect and mood within the previously developed Emotion Ontology, and that of psychiatric disorders in the Mental Disease Ontology. This effort further draws on developments in formal ontology regarding the distinction between normal and abnormal in order to formalise the interconnections. This operational semantic framework is relevant for applications including clarifying psychiatric diagnostic categories, clinical information systems, and the integration and translation of research results across disciplines.
- Perspectives from the NanoSafety Modelling Cluster on the validation criteria for (Q)SAR models used in nanotechnologyTomasz Puzyn, Nina Jeliazkova, Haralambos Sarimveis, Richard L. Marchese Robinson, Vladimir Lobaskin, Robert Rallo, Andrea-N. Richarz, Agnieszka Gajewicz, Manthos G. Papadopulos, Janna Hastings, and 3 more authorsFood and Chemical Toxicology, Feb 2018
Nanotechnology and the production of nanomaterials have been expanding rapidly in recent years. Since many types of engineered nanoparticles are suspected to be toxic to living organisms and to have a negative impact on the environment, the process of designing new nanoparticles and their applications must be accompanied by a thorough risk analysis. (Quantitative) Structure-Activity Relationship ([Q]SAR) modelling creates promising options among the available methods for the risk assessment. These in silico models can be used to predict a variety of properties, including the toxicity of newly designed nanoparticles. However, (Q)SAR models must be appropriately validated to ensure the clarity, consistency and reliability of predictions. This paper is a joint initiative from recently completed European research projects focused on developing (Q)SAR methodology for nanomaterials. The aim was to interpret and expand the guidance for the well-known “OECD Principles for the Validation, for Regulatory Purposes, of (Q)SAR Models”, with reference to nano-(Q)SAR, and present our opinions on the criteria to be fulfilled for models developed for nanoparticles.
2017
- Primer on OntologiesJanna HastingsIn The Gene Ontology Handbook, Feb 2017
As molecular biology has increasingly become a data-intensive discipline, ontologies have emerged as an essential computational tool to assist in the organisation, description and analysis of data. Ontologies describe and classify the entities of interest in a scientific domain in a computationally accessible fashion such that algorithms and tools can be developed around them. The technology that underlies ontologies has its roots in logic-based artificial intelligence, allowing for sophisticated automated inference and error detection. This chapter presents a general introduction to modern computational ontologies as they are used in biology.
- WormJam: A consensus \textitC. elegans Metabolic Reconstruction and Metabolomics Community and Workshop SeriesJanna Hastings, Abraham Mains, Marta Artal-Sanz, Sven Bergmann, Bart P. Braeckman, Jake Bundy, Filipe Cabreiro, Paul Dobson, Paul Ebert, Jake Hattwell, and 28 more authorsWorm, Feb 2017
- Identifiers for the 21st century: How to design, provision, and reuse persistent identifiers to maximize utility and impact of life science data.Julie A McMurry, Nick Juty, Niklas Blomberg, Tony Burdett, Tom Conlin, Nathalie Conte, Mélanie Courtot, John Deck, Michel Dumontier, Donal K Fellows, and 34 more authorsPLoS Biology, Jun 2017
In many disciplines, data are highly decentralized across thousands of online databases (repositories, registries, and knowledgebases). Wringing value from such databases depends on the discipline of data science and on the humble bricks and mortar that make integration possible; identifiers are a core component of this integration infrastructure. Drawing on our experience and on work by other groups, we outline 10 lessons we have learned about the identifier qualities and best practices that facilitate large-scale data integration. Specifically, we propose actions that identifier practitioners (database providers) should take in the design, provision and reuse of identifiers. We also outline the important considerations for those referencing identifiers in various circumstances, including by authors and data generators. While the importance and relevance of each lesson will vary by context, there is a need for increased awareness about how to avoid and manage common identifier problems, especially those related to persistence and web-accessibility/resolvability. We focus strongly on web-based identifiers in the life sciences; however, the principles are broadly relevant to other disciplines.
2016
- ChEBI in 2016: Improved services and an expanding collection of metabolites.Janna Hastings, Gareth Owen, Adriano Dekker, Marcus Ennis, Namrata Kale, Venkatesh Muthukrishnan, Steve Turner, Neil Swainston, Pedro Mendes, and Christoph SteinbeckNucleic Acids Research, Jan 2016
ChEBI is a database and ontology containing information about chemical entities of biological interest. It currently includes over 46,000 entries, each of which is classified within the ontology and assigned multiple annotations including (where relevant) a chemical structure, database cross-references, synonyms and literature citations. All content is freely available and can be accessed online at http://www.ebi.ac.uk/chebi. In this update paper, we describe recent improvements and additions to the ChEBI offering. We have substantially extended our collection of endogenous metabolites for several organisms including human, mouse, Escherichia coli and yeast. Our front-end has also been reworked and updated, improving the user experience, removing our dependency on Java applets in favour of embedded JavaScript components and moving from a monthly release update to a ’live’ website. Programmatic access has been improved by the introduction of a library, libChEBI, in Java, Python and Matlab. Furthermore, we have added two new tools, namely an analysis tool, BiNChE, and a query tool for the ontology, OntoQuery.
- ClassyFire: automated chemical classification with a comprehensive, computable taxonomyYannick Djoumbou Feunang, Roman Eisner, Craig Knox, Leonid Chepelev, Janna Hastings, Gareth Owen, Eoin Fahy, Christoph Steinbeck, Shankar Subramanian, Evan Bolton, and 2 more authorsJournal of Cheminformatics, Dec 2016
Background: Scientists have long been driven by the desire to describe, organize, classify, and compare objects using taxonomies and/or ontologies. In contrast to biology, geology, and many other scientific disciplines, the world of chemistry still lacks a standardized chemical ontology or taxonomy. Several attempts at chemical classification have been made; but they have mostly been limited to either manual, or semi-automated proof-of-principle applications. This is regrettable as comprehensive chemical classification and description tools could not only improve our understanding of chemistry but also improve the linkage between chemistry and many other fields. For instance, the chemical classification of a compound could help predict its metabolic fate in humans, its druggability or potential hazards associated with it, among others. However, the sheer number (tens of millions of compounds) and complexity of chemical structures is such that any manual classification effort would prove to be near impossible. Results: We have developed a comprehensive, flexible, and computable, purely structure-based chemical taxonomy (ChemOnt), along with a computer program (ClassyFire) that uses only chemical structures and structural features to automatically assign all known chemical compounds to a taxonomy consisting of \textgreater4800 different categories. This new chemical taxonomy consists of up to 11 different levels (Kingdom, SuperClass, Class, SubClass, etc.) with each of the categories defined by unambiguous, computable structural rules. Furthermore each category is named using a consensus-based nomenclature and described (in English) based on the characteristic common structural properties of the compounds it contains. The ClassyFire webserver is freely accessible at http://classyfire.wishartlab.com/. Moreover, a Ruby API version is available at https://bitbucket.org/wishartlab/classyfire_api, which provides programmatic access to the ClassyFire server and database. ClassyFire has been used to annotate over 77 million compounds and has already been integrated into other software packages to automatically generate textual descriptions for, and/ or infer biological properties of over 100,000 compounds. Additional examples and applications are provided in this paper. Conclusion: ClassyFire, in combination with ChemOnt (ClassyFire’s comprehensive chemical taxonomy), now allows chemists and cheminformaticians to perform large-scale, rapid and automated chemical classification. Moreover, a freely accessible API allows easy access to more than 77 million “ClassyFire” classified compounds. The results can be used to help annotate well studied, as well as lesser-known compounds. In addition, these chemical classifications can be used as input for data integration, and many other cheminformatics-related tasks.
- Harmonising and linking biomedical and clinical data across disparate data archives to enable integrative cross-biobank researchOla Spjuth, Maria Krestyaninova, Janna Hastings, Huei-Yi Shen, Jani Heikkinen, Melanie Waldenberger, Arnulf Langhammer, Claes Ladenvall, Tõnu Esko, Mats-Åke Persson, and 26 more authorsEuropean Journal of Human Genetics, Apr 2016
A wealth of biospecimen samples are stored in modern globally distributed biobanks. Biomedical researchers worldwide need to be able to combine the available resources to improve the power of large-scale studies. A prerequisite for this effort is to be able to search and access phenotypic, clinical and other information about samples that are currently stored at biobanks in an integrated manner. However, privacy issues together with heterogeneous information systems and the lack of agreed-upon vocabularies have made specimen searching across multiple biobanks extremely challenging. We describe three case studies where we have linked samples and sample descriptions in order to facilitate global searching of available samples for research. The use cases include the ENGAGE (European Network for Genetic and Genomic Epidemiology) consortium comprising at least 39 cohorts, the SUMMIT (surrogate markers for micro- and macro-vascular hard endpoints for innovative diabetes tools) consortium and a pilot for data integration between a Swedish clinical health registry and a biobank. We used the Sample avAILability (SAIL) method for data linking: first, created harmonised variables and then annotated and made searchable information on the number of specimens available in individual biobanks for various phenotypic categories. By operating on this categorised availability data we sidestep many obstacles related to privacy that arise when handling real values and show that harmonised and annotated records about data availability across disparate biomedical archives provide a key methodological advance in pre-analysis exchange of information between biobanks, that is, during the project planning phase.
- Digital Scholarship and Open Science in Psychology and the Behavioral Sciences (Dagstuhl Perpectives Workshop 15302)Alexander Garcia Castro, Janna Hastings, Robert Stevens, and Erich WeichselgartnerDagstuhl Reports, Apr 2016
- libChEBI: an API for accessing the ChEBI databaseNeil Swainston, Janna Hastings, Adriano Dekker, Venkatesh Muthukrishnan, John May, Christoph Steinbeck, and Pedro MendesJournal of Cheminformatics, Mar 2016
ChEBI is a database and ontology of chemical entities of biological interest. It is widely used as a source of identifiers to facilitate unambiguous reference to chemical entities within biological models, databases, ontologies and literature. ChEBI contains a wealth of chemical data, covering over 46,500 distinct chemical entities, and related data such as chemical formula, charge, molecular mass, structure, synonyms and links to external databases. Furthermore, ChEBI is an ontology, and thus provides meaningful links between chemical entities. Unlike many other resources, ChEBI is fully human-curated, providing a reliable, non-redundant collection of chemical entities and related data. While ChEBI is supported by a web service for programmatic access and a number of download files, it does not have an API library to facilitate the use of ChEBI and its data in cheminformatics software.
2015
- BiNChE: a web tool and library for chemical enrichment analysis based on the ChEBI ontology.Pablo Moreno, Stephan Beisken, Bhavana Harsha, Venkatesh Muthukrishnan, Ilinca Tudose, Adriano Dekker, Stefanie Dornfeldt, Franziska Taruttis, Ivo Grosse, Janna Hastings, and 2 more authorsBMC Bioinformatics, Mar 2015
Ontology-based enrichment analysis aids in the interpretation and understanding of large-scale biological data. Ontologies are hierarchies of biologically relevant groupings. Using ontology annotations, which link ontology classes to biological entities, enrichment analysis methods assess whether there is a significant over or under representation of entities for ontology classes. While many tools exist that run enrichment analysis for protein sets annotated with the Gene Ontology, there are only a few that can be used for small molecules enrichment analysis.We describe BiNChE, an enrichment analysis tool for small molecules based on the ChEBI Ontology. BiNChE displays an interactive graph that can be exported as a high-resolution image or in network formats. The tool provides plain, weighted and fragment analysis based on either the ChEBI Role Ontology or the ChEBI Structural Ontology.BiNChE aids in the exploration of large sets of small molecules produced within Metabolomics or other Systems Biology research contexts. The open-source tool provides easy and highly interactive web access to enrichment analysis with the ChEBI ontology tool and is additionally available as a standalone library.
- DINTO: Using OWL Ontologies and SWRL Rules to Infer Drug-Drug Interactions and Their Mechanisms.María Herrero-Zazo, Isabel Segura-Bedmar, Janna Hastings, and Paloma MartínezJournal of chemical information and modeling, Aug 2015
The early detection of drug-drug interactions (DDIs) is limited by the diffuse spread of DDI information in heterogeneous sources. Computational methods promise to play a key role in the identification and explanation of DDIs on a large scale. However, such methods rely on the availability of computable representations describing the relevant domain knowledge. Current modeling efforts have focused on partial and shallow representations of the DDI domain, failing to adequately support computational inference and discovery applications. In this paper, we describe a comprehensive ontology for DDI knowledge (DINTO), which is the first formal representation of different types of DDIs and their mechanisms and its application in the prediction of DDIs. This project has been developed using currently available semantic web technologies, standards, and tools, and we have demonstrated that the combination of drug-related facts in DINTO and Semantic Web Rule Language (SWRL) rules can be used to infer DDIs and their different mechanisms on a large scale. The ontology is available from https://code.google.com/p/dinto/.
- Data standards can boost metabolomics research, and if there is a will, there is a wayPhilippe Rocca-Serra, Reza M. Salek, Masanori Arita, Elon Correa, Saravanan Dayalan, Alejandra Gonzalez-Beltran, Tim Ebbels, Royston Goodacre, Janna Hastings, Kenneth Haug, and 9 more authorsMetabolomics, Nov 2015
Thousands of articles using metabolomics approaches are published every year. With the increasing amounts of data being produced, mere description of investigations as text in manuscripts is not sufficient to enable re-use anymore: the underlying data needs to be published together with the findings in the literature to maximise the benefit from public and private expenditure and to take advantage of an enormous opportunity to improve scientific reproducibility in metabolomics and cognate disciplines. Reporting recommendations in metabolomics started to emerge about a decade ago and were mostly concerned with inventories of the information that had to be reported in the literature for consistency. In recent years, metabolomics data standards have developed extensively, to include the primary research data, derived results and the experimental description and importantly the metadata in a machine-readable way. This includes vendor independent data standards such as mzML for mass spectrometry and nmrML for NMR raw data that have both enabled the development of advanced data processing algorithms by the scientific community. Standards such as ISA-Tab cover essential metadata, including the experimental design, the applied protocols, association between samples, data files and the experimental factors for further statistical analysis. Altogether, they pave the way for both reproducible research and data reuse, including meta-analyses. Further incentives to prepare standards compliant data sets include new opportunities to publish data sets, but also require a little “arm twisting” in the author guidelines of scientific journals to submit the data sets to public repositories such as the NIH Metabolomics Workbench or MetaboLights at EMBL-EBI. In the present article, we look at standards for data sharing, investigate their impact in metabolomics and give suggestions to improve their adoption.
- eNanoMapper: harnessing ontologies to enable data integration for nanomaterial risk assessmentJanna Hastings, Nina Jeliazkova, Gareth Owen, Georgia Tsiliki, Cristian R. Munteanu, Christoph Steinbeck, and Egon WillighagenJournal of Biomedical Semantics, Mar 2015
Engineered nanomaterials (ENMs) are being developed to meet specific application needs in diverse domains across the engineering and biomedical sciences (e.g. drug delivery). However, accompanying the exciting proliferation of novel nanomaterials is a challenging race to understand and predict their possibly detrimental effects on human health and the environment. The eNanoMapper project (www.enanomapper.net) is creating a pan-European computational infrastructure for toxicological data management for ENMs, based on semantic web standards and ontologies. Here, we describe the development of the eNanoMapper ontology based on adopting and extending existing ontologies of relevance for the nanosafety domain. The resulting eNanoMapper ontology is available at http://purl.enanomapper.net/onto/enanomapper.owl. We aim to make the re-use of external ontology content seamless and thus we have developed a library to automate the extraction of subsets of ontology content and the assembly of the subsets into an integrated whole. The library is available (open source) at http://github.com/enanomapper/slimmer/. Finally, we give a comprehensive survey of the domain content and identify gap areas. ENM safety is at the boundary between engineering and the life sciences, and at the boundary between molecular granularity and bulk granularity. This creates challenges for the definition of key entities in the domain, which we also discuss.
- The eNanoMapper database for nanomaterial safety informationNina Jeliazkova, Charalampos Chomenidis, Philip Doganis, Bengt Fadeel, Roland Grafström, Barry Hardy, Janna Hastings, Markus Hegi, Vedrin Jeliazkov, Nikolay Kochev, and 8 more authorsBeilstein Journal of Nanotechnology, Jul 2015
Background: The NanoSafety Cluster, a cluster of projects funded by the European Commision, identified the need for a computational infrastructure for toxicological data management of engineered nanomaterials (ENMs). Ontologies, open standards, and interoperable designs were envisioned to empower a harmonized approach to European research in nanotechnology. This setting provides a number of opportunities and challenges in the representation of nanomaterials data and the integration of ENM information originating from diverse systems. Within this cluster, eNanoMapper works towards supporting the collaborative safety assessment for ENMs by creating a modular and extensible infrastructure for data sharing, data analysis, and building computational toxicology models for ENMs.
- Application of Domain Ontologies to Natural Language Processing: A Case Study for Drug-Drug InteractionsMaría Herrero-Zazo, Isabel Segura-Bedmar, Janna Hastings, and Paloma MartínezInternational Journal of Information Retrieval Research (IJIRR), Jul 2015
Natural Language Processing (NLP) techniques can provide an interesting way to mine the growing biomedical literature, and a promising approach for new knowledge discovery. However, the major bottleneck in this area is that these systems rely on specific resources providing the domain knowledge. Dom...
2014
- The first eNanoMapper prototype: a substance database to support safe-by-designNina Jeliazkova, Philip Doganis, Bengt Fadeel, Roland Grafström, Janna Hastings, Vedrin Jeliazkov, Pekka Kohonen, Cristian R. Munteanu, Haralambos Sarimveis, Bart Smeets, and 3 more authorsIn Proceedings of the 2014 Workshop on Nanoinformatics for Environmental Health and Biomedicine, Belfast, UK, Jul 2014
- Ten recommendations for software engineering in researchJanna Hastings, Kenneth Haug, and Christoph SteinbeckGigaScience, Dec 2014
Abstract. Research in the context of data-driven science requires a backbone of well-written software, but scientific researchers are typically not trained at l
- An ontology for drug-drug interactionsMaría Herrero Zazo, Janna Hastings, Isabel Segura-Bedmar, Samuel Croset, Paloma Martínez Fernández, and Christoph SteinbeckJan 2014
Drug-drug interactions form a significant risk group for adverse effects associ-ated with pharmaceutical treatment. These interactions are often reported in the literature, however, they are sparsely represented in machine-readable re-sources, such as online databases, thesauri or ontologies. These knowledge sources play a pivotal role in Natural Language Processing (NLP) systems since they provide a knowledge representation about the world or a particular do-main. While ontologies for drugs and their effects have proliferated in recent years, there is no ontology capable of describing and categorizing drug-drug in-teractions. Moreover, there is no artifact that represents all the possible mecha-nisms that can lead to a DDI. To fill this gap we propose DINTO, an ontology for drug-drug interactions and their mechanisms. In this paper we describe the classes, relationships and overall structure of DINTO. The ontology is free for use and available at https://code.google.com/p/dinto/
- Interdisciplinary perspectives on the development, integration, and application of cognitive ontologiesJanna Hastings, Gwen A. Frishkoff, Barry Smith, Mark Jensen, Russell A. Poldrack, Jane Lomax, Anita Bandrowski, Fahim Imam, Jessica A. Turner, and Maryann E. MartoneFrontiers in Neuroinformatics, Jan 2014
We discuss recent progress in the development of cognitive ontologies and summarize three challenges in the coordinated development and application of these resources. Challenge 1 is to adopt a standardized definition for cognitive processes. We describe three possibilities and recommend one that is consistent with the standard view in cognitive and biomedical sciences. Challenge 2 is harmonization. Gaps and conflicts in representation must be resolved so that these resources can be combined for mark-up and interpretation of multi-modal data. Finally, Challenge 3 is to test the utility of these resources for large-scale annotation of data, search and query, and knowledge discovery and integration. As term definitions are tested and revised, harmonization should enable coordinated updates across ontologies. However, the true test of these definitions will be in their community-wide adoption which will test whether they support valid inferences about psychological and neuroscientific data.
- Evaluating the Emotion Ontology through use in the self-reporting of emotional responses at an academic conferenceJanna Hastings, Andy Brass, Colin Caine, Caroline Jay, and Robert StevensJournal of Biomedical Semantics, Sep 2014
We evaluate the application of the Emotion Ontology (EM) to the task of self-reporting of emotional experience in the context of audience response to academic presentations at the International Conference on Biomedical Ontology (ICBO). Ontology evaluation is regarded as a difficult task. Types of ontology evaluation range from gauging adherence to some philosophical principles, following some engineering method, to assessing fitness for purpose. The Emotion Ontology (EM) represents emotions and all related affective phenomena, and should enable self-reporting or articulation of emotional states and responses; how do we know if this is the case? Here we use the EM ‘in the wild’ in order to evaluate the EM’s ability to capture people’s self-reported emotional responses to a situation through use of the vocabulary provided by the EM.
2013
- Dovetailing biology and chemistry: integrating the Gene Ontology with the ChEBI chemical ontology.David P. Hill, Nico Adams, Mike Bada, Colin Batchelor, Tanya Z. Berardini, Heiko Dietze, Harold J. Drabkin, Marcus Ennis, Rebecca E. Foulger, Midori A. Harris, and 9 more authorsBMC genomics, Jul 2013
The Gene Ontology (GO) facilitates the description of the action of gene products in a biological context. Many GO terms refer to chemical entities that participate in biological processes. To facilitate accurate and consistent systems-wide biological representation, it is necessary to integrate the chemical view of these entities with the biological view of GO functions and processes. We describe a collaborative effort between the GO and the Chemical Entities of Biological Interest (ChEBI) ontology developers to ensure that the representation of chemicals in the GO is both internally consistent and in alignment with the chemical expertise captured in ChEBI. We have examined and integrated the ChEBI structural hierarchy into the GO resource through computationally-assisted manual curation of both GO and ChEBI. Our work has resulted in the creation of computable definitions of GO terms that contain fully defined semantic relationships to corresponding chemical terms in ChEBI. The set of logical definitions using both the GO and ChEBI has already been used to automate aspects of GO development and has the potential to allow the integration of data across the domains of biology and chemistry. These logical definitions are available as an extended version of the ontology from http://purl.obolibrary.org/obo/go/extensions/go-plus.owl.
- The ChEBI reference database and ontology for biologically relevant chemistry: enhancements for 2013Janna Hastings, Paula de Matos, Adriano Dekker, Marcus Ennis, Bhavana Harsha, Namrata Kale, Venkatesh Muthukrishnan, Gareth Owen, Steve Turner, Mark Williams, and 1 more authorNucleic Acids Research, Jul 2013
- MetaboLights–an open-access general-purpose repository for metabolomics studies and associated meta-dataKenneth Haug, Reza M. Salek, Pablo Conesa, Janna Hastings, Paula Matos, Mark Rijnbeek, Tejasvi Mahendraker, Mark Williams, Steffen Neumann, Philippe Rocca-Serra, and 5 more authorsNucleic Acids Research, Jul 2013
- Chemical Ontologies for Standardization, Knowledge Discovery, and Data MiningJanna Hastings and Christoph SteinbeckIn Data Mining in Drug Discovery, Jul 2013
This chapter contains sections titled: Introduction Background Chemical Ontologies Standardization Knowledge Discovery Data Mining Conclusions
- Shape Perception in ChemistryJanna Hastings, Colin Batchelor, and Mitsuhiro OkadaProceedings of SHAPES, Jul 2013
Organic chemists make extensive use of a diagrammatic language for designing, exchanging and analysing the features of chemicals. In this language, chemicals are represented on a flat (2D) plane following standard stylistic conventions. In the search for novel drugs and therapeutic agents, vast quantities of chemical data are generated and subjected to virtual screening procedures that harness algorithmic features and complex statistical models. However, in silico approaches do not yet compare to the abilities of experienced chemists in detecting more subtle features relevant for evaluating how likely a molecule is to be suitable to a given purpose. Our hypothesis is that one reason for this discrepancy is that human perceptual capabilities, particularly that of ‘gestalt’ shape perception, make additional information available to our reasoning processes that are not available to in silico processes. This contribution investigates this hypothesis.
- The MetaboLights repository: curation challenges in metabolomicsReza M. Salek, Kenneth Haug, Pablo Conesa, Janna Hastings, Mark Williams, Tejasvi Mahendraker, Eamonn Maguire, Alejandra N. González-Beltrán, Philippe Rocca-Serra, Susanna-Assunta Sansone, and 1 more authorDatabase, Jan 2013
Abstract. MetaboLights is the first general-purpose open-access curated repository for metabolomic studies, their raw experimental data and associated metadata,
- UniChem: a unified chemical structure cross-referencing and identifier tracking systemJon Chambers, Mark Davies, Anna Gaulton, Anne Hersey, Sameer Velankar, Robert Petryszak, Janna Hastings, Louisa Bellis, Shaun McGlinchey, and John P. OveringtonJournal of Cheminformatics, Jan 2013
UniChem is a freely available compound identifier mapping service on the internet, designed to optimize the efficiency with which structure-based hyperlinks may be built and maintained between chemistry-based resources. In the past, the creation and maintenance of such links at EMBL-EBI, where several chemistry-based resources exist, has required independent efforts by each of the separate teams. These efforts were complicated by the different data models, release schedules, and differing business rules for compound normalization and identifier nomenclature that exist across the organization. UniChem, a large-scale, non-redundant database of Standard InChIs with pointers between these structures and chemical identifiers from all the separate chemistry resources, was developed as a means of efficiently sharing the maintenance overhead of creating these links. Thus, for each source represented in UniChem, all links to and from all other sources are automatically calculated and immediately available for all to use. Updated mappings are immediately available upon loading of new data releases from the sources. Web services in UniChem provide users with a single simple automatable mechanism for maintaining all links from their resource to all other sources represented in UniChem. In addition, functionality to track changes in identifier usage allows users to monitor which identifiers are current, and which are obsolete. Lastly, UniChem has been deliberately designed to allow additional resources to be included with minimal effort. Indeed, the recent inclusion of data sources external to EMBL-EBI has provided a simple means of providing users with an even wider selection of resources with which to link to, all at no extra cost, while at the same time providing a simple mechanism for external resources to link to all EMBL-EBI chemistry resources.
2012
- Modelling Highly Symmetrical Molecules: Linking Ontologies and GraphsOliver Kutz, Janna Hastings, and Till MossakowskiIn Artificial Intelligence: Methodology, Systems, and Applications, Jan 2012
Methods for automated classification of chemical data depend on identifying interesting parts and properties. However, classes of chemical entities which are highly symmetrical and contain large numbers of homogeneous parts (such as carbon atoms) are not straightforwardly classified in this fashion. One such class of molecules is the fullerene family, which shows potential for many novel applications including in biomedicine. The Web Ontology Language OWL cannot be used to represent the structure of fullerenes, as their structure is not treeshaped. While individual members of the fullerene class can be modelled in standard FOL, expressing the properties of the class as a whole (independent of the count of atoms of the members) requires second-order quantification. Given the size of chemical ontologies such as ChEBI, using second-order expressivity in the general case is prohibitively expensive to practical applications. To address these conflicting requirements, we introduce a novel framework in which we heterogeneously integrate standard ontological modelling with monadic second-order reasoning over chemical graphs, enabling various kinds of information flow between the distinct representational layers.
- Ontologies for Human Behavior Analysis and Their Application to Clinical DataJanna Hastings and Stefan SchulzIn International Review of Neurobiology, Jan 2012
Mental and behavioral disorders are common in all countries and represent a significant portion of the public health burden in developed nations. The human cost of these disorders is immense, yet treatment options for sufferers are currently limited, with many patients failing to respond sufficiently to currently available interventions.
- Structure-based classification and ontology in chemistry.Janna Hastings, Despoina Magka, Colin Batchelor, Lian Duan, Robert Stevens, Marcus Ennis, and Christoph SteinbeckJournal of cheminformatics, Apr 2012
Recent years have seen an explosion in the availability of data in the chemistry domain. With this information explosion, however, retrieving relevant results from the available information, and organising those results, become even harder problems. Computational processing is essential to filter and organise the available resources so as to better facilitate the work of scientists. Ontologies encode expert domain knowledge in a hierarchically organised machine-processable format. One such ontology for the chemical domain is ChEBI. ChEBI provides a classification of chemicals based on their structural features and a role or activity-based classification. An example of a structure-based class is ’pentacyclic compound’ (compounds containing five-ring structures), while an example of a role-based class is ’analgesic’, since many different chemicals can act as analgesics without sharing structural features. Structure-based classification in chemistry exploits elegant regularities and symmetries in the underlying chemical domain. As yet, there has been neither a systematic analysis of the types of structural classification in use in chemistry nor a comparison to the capabilities of available technologies. We analyze the different categories of structural classes in chemistry, presenting a list of patterns for features found in class definitions. We compare these patterns of class definition to tools which allow for automation of hierarchy construction within cheminformatics and within logic-based ontology technology, going into detail in the latter case with respect to the expressive capabilities of the Web Ontology Language and recent extensions for modelling structured objects. Finally we discuss the relationships and interactions between cheminformatics approaches and logic-based approaches. Systems that perform intelligent reasoning tasks on chemistry data require a diverse set of underlying computational utilities including algorithmic, statistical and logic-based tools. For the task of automatic structure-based classification of chemical entities, essential to managing the vast swathes of chemical data being brought online, systems which are capable of hybrid reasoning combining several different approaches are crucial. We provide a thorough review of the available tools and methodologies, and identify areas of open research.
- Process attributes in bio-ontologiesAndre Andrade, Ward Blonde, Janna Hastings, and Stefan SchulzBMC Bioinformatics, Apr 2012
BACKGROUND:Biomedical processes can provide essential information about the (mal-) functioning of an organism and are thus frequently represented in biomedical terminologies and ontologies, including the GO Biological Process branch. These processes often need to be described and categorised in terms of their attributes, such as rates or regularities. The adequate representation of such process attributes has been a contentious issue in bio-ontologies recently; and domain ontologies have correspondingly developed ad hoc workarounds that compromise interoperability and logical consistency.RESULTS:We present a design pattern for the representation of process attributes that is compatible with upper ontology frameworks such as BFO and BioTop. Our solution rests on two key tenets: firstly, that many of the sorts of process attributes which are biomedically interesting can be characterised by the ways that repeated parts of such processes constitute, in combination, an overall process; secondly, that entities for which a full logical definition can be assigned do not need to be treated as primitive within a formal ontology framework. We apply this approach to the challenge of modelling and automatically classifying examples of normal and abnormal rates and patterns of heart beating processes, and discuss the expressivity required in the underlying ontology representation language. We provide full definitions for process attributes at increasing levels of domain complexity.CONCLUSIONS:We show that a logical definition of process attributes is feasible, though limited by the expressivity of DL languages so that the creation of primitives is still necessary. This finding may endorse current formal upper-ontology frameworks as a way of ensuring consistency, interoperability and clarity.
- Self-organizing ontology of biochemically relevant small moleculesLeonid L. Chepelev, Janna Hastings, Marcus Ennis, Christoph Steinbeck, and Michel DumontierBMC Bioinformatics, Jan 2012
The advent of high-throughput experimentation in biochemistry has led to the generation of vast amounts of chemical data, necessitating the development of novel analysis, characterization, and cataloguing techniques and tools. Recently, a movement to publically release such data has advanced biochemical structure-activity relationship research, while providing new challenges, the biggest being the curation, annotation, and classification of this information to facilitate useful biochemical pattern analysis. Unfortunately, the human resources currently employed by the organizations supporting these efforts (e.g. ChEBI) are expanding linearly, while new useful scientific information is being released in a seemingly exponential fashion. Compounding this, currently existing chemical classification and annotation systems are not amenable to automated classification, formal and transparent chemical class definition axiomatization, facile class redefinition, or novel class integration, thus further limiting chemical ontology growth by necessitating human involvement in curation. Clearly, there is a need for the automation of this process, especially for novel chemical entities of biological interest.
- Toxicology ontology perspectivesBarry Hardy, Gordana Apic, Philip Carthew, Dominic Clark, David Cook, Ian Dix, Sylvia Escher, Janna Hastings, David J. Heard, Nina Jeliazkova, and 11 more authorsALTEX - Alternatives to animal experimentation, May 2012
- A toxicology ontology roadmapBarry Hardy, Gordana Apic, Philip Carthew, Dominic Clark, David Cook, Ian Dix, Sylvia Escher, Janna Hastings, David J. Heard, Nina Jeliazkova, and 11 more authorsALTEX - Alternatives to animal experimentation, May 2012
- Accessing and Using Chemical Property DatabasesJanna Hastings, Zara Josephs, and Christoph SteinbeckIn Computational Toxicology: Volume I, May 2012
Chemical compounds participate in all the processes of life. Understanding the complex interactions of small molecules such as metabolites and drugs and the biological macromolecules that consume and produce them is key to gaining a wider understanding in a systemic context. Chemical property databases collect information on the biological effects and physicochemical properties of chemical entities. Accessing and using such databases is key to understanding the chemistry of toxic molecules. In this chapter, we present methods to search, understand, download, and manipulate the wealth of information available in public chemical property databases, with particular focus on the database of Chemical Entities of Biological Interest (ChEBI).
- Requirements for Semantic BiobanksAndré Q Andrade, Markus Kreuzthaler, Janna Hastings, Maria Krestyaninova, and Stefan SchulzStud Health Technol Inform, May 2012
World-wide availability of biobank samples is a great desideratum for biomedical researchers. We describe the use case of biobank information retrieval that requires the semantic descriptions of biobank samples and of clinical information. In addition we sketch the foundations of an ontology for biobanks, as a basis on which distributed biobank indexing and retrieval systems can be built. We advocate that a detailed and robust representation of this kind of information improves and allows complex queries that will certainly arise to explore the full potential of biobanks.
- Structured chemical class definitions and automated matching for chemical ontology evolutionLian Duan, Janna Hastings, Paula Matos, Marcus Ennis, and Christoph SteinbeckJournal of Cheminformatics, May 2012
- CheminformaticsJoerg Kurt Wegner, Aaron Sterling, Rajarshi Guha, Andreas Bender, Jean-Loup Faulon, Janna Hastings, Noel O’Boyle, John Overington, Herman Van Vlijmen, and Egon WillighagenCommunications of the ACM, Nov 2012
Open-source chemistry software and molecular databases broaden the research horizons of drug discovery.
- A Database for Chemical Proteomics: ChEBIPaula Matos, Nico Adams, Janna Hastings, Pablo Moreno, and Christoph SteinbeckIn Chemical Proteomics: Methods and Protocols, Nov 2012
Chemical proteomics is concerned with the identification of protein targets interacting with small molecules. Hence, the availability of a high quality and free resource storing small molecules is essential for the future development of the field. The Chemical Entities of Biological Interest (ChEBI) database is one such database. The scope of ChEBI includes any constitutionally or isotopically distinct atom, molecule, ion, ion pair, radical, radical ion, complex, conformer, etc., identifiable as a separately distinguishable entity. These entities in question are either products of nature or synthetic products used to intervene in the processes of living organisms. In addition, ChEBI contains a chemical ontology which relates the small molecules with each other thereby making it easier for users to discover data. The ontology also describes the biological roles that the small molecules are active in. The ChEBI database also provides a central reference point in which to access a variety of bioinformatics data points such as pathways and their biochemical reactions; expression data; protein sequence and structures.
- Annotating affective neuroscience data with the Emotion OntologyJanna Hastings, Werner Ceusters, Kevin Mulligan, and Barry SmithIn Proceedings of the 3rd International Conference on Biomedical Ontologies, Workshop Towards an Ontology of Mental Functioning., Nov 2012
The Emotion Ontology is an ontology covering all aspects of emotional and affective mental functioning. It is being developed following the principles of the OBO Foundry and Ontological Realism. This means ...
2011
- Modularization requirements in bio-ontologies: A case study of ChEBIJanna Hastings, Colin Batchelor, Stefan Schulz, and Christoph SteinbeckIn Workshop on Modular Ontologies (WoMO 2011), Ljubljana, Slovenia, Frontiers in Artificial Intelligence and Applications, Nov 2011
- The Chemical Information Ontology: Provenance and Disambiguation for Chemical Data on the Biological Semantic WebJanna Hastings, Leonid Chepelev, Egon Willighagen, Nico Adams, Christoph Steinbeck, and Michel DumontierPLoS ONE, Nov 2011
\textlessp\textgreaterCheminformatics is the application of informatics techniques to solve chemical problems \textlessitalic\textgreaterin silico\textless/italic\textgreater. There are many areas in biology where cheminformatics plays an important role in computational research, including metabolism, proteomics, and systems biology. One critical aspect in the application of cheminformatics in these fields is the accurate exchange of data, which is increasingly accomplished through the use of ontologies. Ontologies are formal representations of objects and their properties using a logic-based ontology language. Many such ontologies are currently being developed to represent objects across all the domains of science. Ontologies enable the definition, classification, and support for querying objects in a particular domain, enabling intelligent computer applications to be built which support the work of scientists both within the domain of interest and across interrelated neighbouring domains. Modern chemical research relies on computational techniques to filter and organise data to maximise research productivity. The objects which are manipulated in these algorithms and procedures, as well as the algorithms and procedures themselves, enjoy a kind of virtual life within computers. We will call these \textlessitalic\textgreaterinformation entities\textless/italic\textgreater. Here, we describe our work in developing an ontology of chemical information entities, with a primary focus on data-driven research and the integration of calculated properties (descriptors) of chemical entities within a semantic web context. Our ontology distinguishes algorithmic, or procedural information from declarative, or factual information, and renders of particular importance the annotation of provenance to calculated data. The Chemical Information Ontology is being developed as an open collaborative project. More details, together with a downloadable OWL file, are available at \textlessext-link xmlns:xlink="http://www.w3.org/1999/xlink" ext-link-type="uri" xlink:href="http://code.google.com/p/semanticchemistry/" xlink:type="simple"\textgreaterhttp://code.google.com/p/semanticchemistry/\textless/ext-link\textgreater (license: CC-BY-SA).\textless/p\textgreater
- Hyperontology for the Biomedical Ontologist —A Sketch and Some Examples—Oliver Kutz, Till Mossakowski, Janna Hastings, Alexander Garcia Castro, and Aleksandra SojicIn Proceedings of the International Conference on Biomedical Ontologies 2011, Workshop on Working with Multiple Biomedical Ontologies, Nov 2011
The Hyperontology framework has been recently introduced to provide a general methodology for heterogeneous ontology design, i.e. the construction of ontologies that have parts, or modules, written in different formalisms, and which are interlinked in complex ways. We here present a brief outline of this framework, discuss its features and merits, and illustrate its usefulness for the domain of biomedical ontology design by providing and discussing a number of examples.
2010
- Chemical Entities of Biological Interest: an updatePaula Matos, Rafael Alcántara, Adriano Dekker, Marcus Ennis, Janna Hastings, Kenneth Haug, Inmaculada Spiteri, Steve Turner, and Christoph SteinbeckNucleic Acids Research, Jan 2010
ABSTRACT. Chemical Entities of Biological Interest (ChEBI) is a freely available dictionary of molecular entities focused on ‘small’ chemical compounds. The mol
2009
- Towards automatic classification within the ChEBI ontologyJanna Hastings, Paula Matos, Marcus Ennis, and Christoph SteinbeckJan 2009
- ChEBI: An Open Bioinformatics and Cheminformatics ResourceKirill Degtyarenko, Janna Hastings, Paula de Matos, and Marcus EnnisCurrent Protocols in Bioinformatics, Jan 2009
Chemical Entities of Biological Interest (ChEBI) is a freely available dictionary of molecular entities focused on “small” chemical compounds. This unit provides a detailed guide to browsing, searching, downloading, and programmatic access to the ChEBI database. Curr. Protoc. Bioinform. 26:14.9.1-14.9.20. © 2009 by John Wiley & Sons, Inc.
2008
- ChEBI: a database and ontology for chemical entities of biological interest.Kirill Degtyarenko, Paula Matos, Marcus Ennis, Janna Hastings, Martin Zbinden, Alan McNaught, Rafael Alcántara, Michael Darsow, Mickaël Guedj, and Michael AshburnerNucleic Acids Research, Jan 2008
Chemical Entities of Biological Interest (ChEBI) is a freely available dictionary of molecular entities focused on ’small’ chemical compounds. The molecular entities in question are either natural products or synthetic products used to intervene in the processes of living organisms. Genome-encoded macromolecules (nucleic acids, proteins and peptides derived from proteins by cleavage) are not as a rule included in ChEBI. In addition to molecular entities, ChEBI contains groups (parts of molecular entities) and classes of entities. ChEBI includes an ontological classification, whereby the relationships between molecular entities or classes of entities and their parents and/or children are specified. ChEBI is available online at http://www.ebi.ac.uk/chebi/
















































































